
We use cookies to ensure our website works properly and to personalise your experience. Cookies policy

We use cookies to ensure our website works properly and to personalise your experience. Cookies policy
Department of Pharmaceutical Chemistry, Madras Medical College, Chennai, Tamil Nadu, India 600003
Osteoarthritis (OA) is a degenerative joint disease marked by Articular cartilage breakdown, subchondral bone remodelling, and synovial inflammation. Current treatments focus primarily on symptom relief, highlighting the urgent need for disease-modifying osteoarthritis drugs (DMOADs). This study presents a dual-targeted therapeutic approach by inhibiting two critical enzymes involved in cartilage degradation: Discoidin Domain Receptor 2 (DDR2) and ADAMTS-5 (A Disintegrin and Metalloproteinase with Thrombospondin Motifs 5). A virtual library of 175 quinazolinone-based ligands was developed using pharmacophore modelling. High-resolution protein structures (PDB IDs: 7AZB for DDR2 and 2RJQ for ADAMTS-5) were validated through Ramachandran plot analysis to ensure structural accuracy. Ligand structures underwent energy minimization, and their novelty was verified via the PubChem database. ADMET analysis confirmed favourable pharmacokinetic and toxicity profiles in accordance with Lipinski’s Rule of Five. Molecular docking studies revealed strong binding interactions for several ligands, with HMJ48, HMJ92, and HMJ155 demonstrating superior binding affinities compared to the standard drug Celecoxib. These findings suggest that quinazolinone derivatives hold significant promise as dual inhibitors for OA treatment. The results offer a foundation for future experimental studies aimed at developing effective, disease-modifying therapies.
Osteoarthritis (OA) is the most prevalent chronic joint disorder, characterized by the progressive degeneration of articular cartilage, subchondral bone remodeling, synovial inflammation, and impaired joint function(1). It commonly affects weight-bearing joints such as the knees, hips, spine, and also the hands, leading to pain, stiffness, and reduced mobility. Risk factors include aging, obesity, genetic predisposition, prior joint injuries, and metabolic abnormalities. Globally, OA imposes a significant health burden, particularly among elderly populations, and is recognized as a leading cause of disability(2). Despite considerable advances in OA research, current therapeutic strategies predominantly aim to alleviate symptoms through the use of analgesics, non steroidal anti-inflammatory drugs (NSAIDs), corticosteroids, and lifestyle interventions. While these approaches provide symptomatic relief, they do not halt or reverse disease progression. This has driven intense research efforts toward developing disease-modifying osteoarthritis drugs (DMOADs) that target key molecular pathways involved in cartilage breakdown and inflammation. One such target is Discoidin Domain Receptor 2 (DDR2), a collagen-activated receptor tyrosine kinase implicated in OA pathogenesis (3). DDR2 activation enhances the expression of matrix metalloproteinase (MMPs), which play a crucial role in extracellular matrix (ECM) degradation and cartilage destruction(4). Additionally, enzymes from the A Disintegrin and Metalloproteinase with Thrombospondin Motifs (ADAMTS) family, particularly ADAMTS-4 (aggrecanase-1) and ADAMTS-5 (aggrecanase-2), are key mediators of aggrecan cleavage within the interglobular domain of cartilage tissue. Among these, ADAMTS-5 has emerged as the predominant aggrecanase in OA. Genetic studies have shown that deletion or inhibition of ADAMTS-5 significantly reduces aggrecan degradation and cartilage erosion in OA models. Although both ADAMTS-4 and ADAMTS-5 can cleave aggrecan, ADAMTS-5 is more catalytically active under pathological conditions. Aggrecan is a major structural proteoglycan within the extracellular matrix of articular cartilage, essential for maintaining the tissue's mechanical integrity. Its high negative charge enables water retention, conferring the cartilage with the ability to resist compressive forces. Loss of aggrecan disrupts these properties, accelerating cartilage degeneration (5). Therefore, targeting both DDR2 and ADAMTS-5 offers a promising dual therapeutic strategy, capable of mitigating cartilage degradation, inflammation, and aberrant bone remodeling associated with OA. In parallel, quinazolinone and its derivatives, a class of heterocyclic compounds, have gained interest due to their broad pharmacological activities, including anti-inflammatory and anticancer effects(6). Their structural versatility allows interaction with a wide range of biological targets, making them suitable scaffolds for the development of dual inhibitors against DDR2 and ADAMTS-5. With the advancement of computational drug discovery techniques, particularly molecular docking, the identification and optimization of potential therapeutic candidates have become significantly more time efficient and cost-effective(7). These in silico methods enable rapid screening of large compound libraries and provide valuable insights into ligand–target interactions at the molecular level. In this study, a library of 175 novel ligands was rationally designed and evaluated for their binding affinity toward both DDR2 and ADAMTS-5, aiming to identify potential dual inhibitors. The findings offer a promising starting point for the development of disease-modifying osteoarthritis drugs (DMOADs) and lay the groundwork for future experimental validation and clinical translation.
MATERIALS AND METHODS
PHARMACOPHORE MODELLING
The Pharmit server was utilized to generate pharmacophore models and screen them against extensive chemical compound libraries, including PubChem, ChEMBL, and the ZINC database. The pharmacophore features were identified using RCSB PDB IDs 7AZB and 2RJQ as input structures(8). These pharmacophoric features were considered critical during ligand design.
RAMACHANDRAN PLOT ANALYSIS
The quality of the crystal structures for 7AZB and 2RJQ was validated using Ramachandran plot analysis(9). Most amino acid residues were located in favored regions, indicating the structural reliability of the receptors and supporting the accuracy of the subsequent docking studies.
LIGAND PREPARATION
A virtual library comprising 175 novel ligands was developed based on the pharmacophore models. These ligands were sketched using ChemSketch software (ACD/Labs, Version 2023 1.0) (10) and saved in MOL format for downstream processes.
Energy Minimization: The ligand structures underwent energy minimization using the Chem3D Ultra software (PerkinElmer Informatics)(11). The MM2 force field was applied, and the minimization was continued until stable, low-energy conformations were obtained.
Novelty Assessment: To ensure chemical novelty, all designed ligands were screened against public chemical databases such as PubChem(12). Ligands not previously reported were selected for further analysis. The energy-minimized structures were converted into PDBQT format using AutoDock Tools 1.5.6, preparing them for molecular docking studies.
RECEPTOR PREPARATION
The target proteins DDR2 and ADAMTS5 were selected based on their high-resolution X-ray crystallographic structures and validated by Ramachandran plot. The target was downloaded from protein data bank(13) (PDB IDs: 7AZB for DDR2 and 2RJQ for ADAMTS5). Receptors were pre-processed by removing water molecules and cofactors, adding polar hydrogens, and assigning Kollman charges using AutoDock Tools 1.5.6.
ADMET PREDICTION
The pharmacokinetic and toxicity profiles of the ligands were predicted using Swiss ADME (14) and OSIRIS Property Explorer(15). Parameters such as drug-likeness, bioavailability, and adherence to Lipinski's Rule of Five were evaluated. OSIRIS provided toxicity assessments for tumorigenicity, mutagenicity, irritancy, and reproductive toxicity. Green indicators denoted non-toxic and safe compounds. Red indicators flagged molecules with potential toxic or adverse effects.
MOLECULAR DOCKING
Molecular docking was conducted using AutoDock Tools 1.5.6(16) to evaluate the binding affinities of the ligands with DDR2 and ADAMTS5. The docking analysis provided insights into the binding orientation and interaction strength, expressed as binding energy values in kcal/mol.
Binding Site Prediction: The CB-DOCK2(17) web server was used to predict the active binding cavities of the target proteins. This structure-based approach identifies ligand-binding pockets by clustering solvent-accessible surface areas, providing information about the coordinates, size, and volume of the binding cavities. This ensured that docking was focused on relevant, functionally important regions.
Visualization Studies: The receptor–ligand interactions were visualized using Molegro Molecular Viewer(18). Key binding residues, interaction types, and distances were analyzed to gain deeper insights into the binding mechanisms of the ligands.
RESULTS AND DISCUSSION
VIRTUAL LIBRARY OF LIGANDS
The pharmacophoric features considered for DDR2 and ADAMTS-5 included hydrophobic moieties, hydrogen bond acceptors, and hydrogen bond donors. These features were essential in guiding the rational design of ligand structures tailored for optimal interaction with the target proteins.
NOVELTY CHECK
Out of the 175 designed ligands, 153 were predicted to be novel based on a comprehensive search against the Pubchem chemical database. The detailed results of the novelty assessment are presented in the supplementary TABLE I.
TABLE NO.I: Table consist of Novel ligands and already existing compounds
|
NOVEL LIGANDS |
ALREADY EXISTING COMPUNDS |
|
H1,H2,H3,H4,H5,H6,H7,H8,H9,H10,H11,H12,H13,H14,H15,H16,H17,H18,H19,H20,H21,H22,H23,H24,H25,H26,H27,H28,H29,H30,H31,H32,H33,H34,H35,H36,H37,H38,H39,H40,H41,H42,H43,H44,H45,H46,H47,H48,H49,H50,H51,H52,H53,H54,H55,H56,H57,H58,H59,H60,H61,H62,H63,H64,H65,H66,H67,H68,H69,H70,H71,H74,H75,H77,H78,H79,H80,H81,H82,H83,H84,H85,H86,H87,H88,H89,H90,H91,H92,H93,H94,H95,H96,H97,H98,H99,H100,H101,H102,H103,H104,H105,H106,H108,H110,H112,H113,H114,H115,H120,H121,H122,H123,H124,H125,H126,H127,H129,H130,H131,H132,H133,H134,H135,H136,H137,H138,H139,H140,H141,H142,H143,H144,H145,H150,H151,H152,H153,H154,H155,H157,H158,H159,H160,H162,H163,H164,H166,H170,H171,H172,H173,H174 |
H72,H73,H76,H107,H109,H111,H116,H117,H118,H119,H128,H146,H147,H148,H149,H156,H161,H165,H175,H167,H168,H169 |
IN-SILICO ADMET STUDIES
The ligands selected for further development were evaluated for their pharmacokinetic properties and were found to comply with Lipinski’s Rule of Five. The molecular weights of these compounds ranged between 237.76 and 456.50 Daltons, while their log P values were within the range of 0.31 to 3.67 comparable to the known reference drugs Celecoxib. Most of the designed ligands are with good drug likeness properties and non-toxic. Data provided in the supplementary TABLE II.
TABLE NO. II: ADMET Properties of Novel ligands
|
Lig no. |
M |
T |
I |
R |
LOG P |
MOLECULAR WEIGHT |
HBD |
HBA |
RULE OF 5 |
|
1 |
NO |
NO |
NO |
NO |
2.57 |
305.33 |
1 |
4 |
0 |
|
2 |
NO |
NO |
NO |
NO |
2.89 |
277.32 |
1 |
3 |
0 |
|
3 |
NO |
NO |
NO |
NO |
2.89 |
297.74 |
1 |
3 |
0 |
|
4 |
NO |
NO |
NO |
NO |
2.16 |
279.29 |
4 |
4 |
0 |
|
5 |
NO |
NO |
NO |
NO |
2.34 |
337.33 |
2 |
5 |
0 |
|
6 |
NO |
NO |
NO |
NO |
2.80 |
321.33 |
1 |
5 |
0 |
|
7 |
NO |
NO |
NO |
NO |
2.52 |
263.29 |
1 |
3 |
0 |
|
8 |
NO |
NO |
NO |
NO |
2.75 |
277.32 |
1 |
3 |
0 |
|
10 |
NO |
NO |
NO |
NO |
2.48 |
278.31 |
2 |
3 |
0 |
|
11 |
NO |
NO |
NO |
NO |
2.69 |
292.34 |
2 |
3 |
0 |
|
12 |
NO |
NO |
NO |
NO |
1.98 |
264.28 |
2 |
3 |
0 |
|
13 |
NO |
NO |
NO |
NO |
2.22 |
293.32 |
3 |
3 |
0 |
|
14 |
NO |
NO |
NO |
NO |
2.55 |
307.35 |
3 |
3 |
0 |
|
15 |
NO |
NO |
NO |
YES |
2.67 |
335.38 |
3 |
4 |
0 |
|
16 |
NO |
NO |
NO |
NO |
3.07 |
313.74 |
1 |
4 |
0 |
|
17 |
NO |
NO |
NO |
NO |
3.03 |
311.77 |
1 |
3 |
0 |
|
18 |
NO |
NO |
NO |
NO |
2.96 |
291.35 |
1 |
3 |
0 |
|
19 |
NO |
YES |
NO |
NO |
1.92 |
279.30 |
3 |
4 |
0 |
|
20 |
NO |
NO |
NO |
NO |
1.98 |
264.28 |
2 |
3 |
0 |
|
21 |
NO |
YES |
NO |
NO |
1.81 |
313.74 |
3 |
4 |
0 |
|
22 |
NO |
YES |
NO |
NO |
2.17 |
293.32 |
3 |
4 |
0 |
|
23 |
NO |
NO |
NO |
NO |
2.21 |
298.73 |
2 |
3 |
0 |
|
24 |
YES |
YES |
NO |
NO |
2.59 |
337.40 |
3 |
3 |
0 |
|
25 |
YES |
NO |
NO |
NO |
2.69 |
322.38 |
2 |
3 |
0 |
|
26 |
YES |
YES |
NO |
NO |
2.47 |
325.39 |
4 |
3 |
0 |
|
27 |
NO |
NO |
NO |
NO |
2.05 |
237.26 |
1 |
4 |
0 |
|
28 |
NO |
NO |
NO |
NO |
1.94 |
265.27 |
1 |
5 |
0 |
|
29 |
NO |
NO |
NO |
NO |
2.39 |
294.31 |
2 |
4 |
0 |
|
30 |
YES |
YES |
NO |
NO |
2.57 |
306.32 |
2 |
4 |
0 |
|
31 |
NO |
NO |
NO |
NO |
3.04 |
306.38 |
2 |
3 |
0 |
|
32 |
NO |
NO |
NO |
NO |
3.29 |
340.81 |
2 |
3 |
0 |
|
33 |
NO |
NO |
NO |
NO |
1.85 |
324.33 |
3 |
4 |
0 |
|
34 |
NO |
NO |
NO |
NO |
2.38 |
294.31 |
3 |
4 |
0 |
|
35 |
NO |
NO |
NO |
NO |
2.11 |
314.73 |
3 |
4 |
0 |
|
36 |
NO |
NO |
NO |
NO |
1.60 |
308.29 |
3 |
5 |
0 |
|
37 |
NO |
NO |
NO |
NO |
2.00 |
342.74 |
3 |
5 |
0 |
|
38 |
NO |
NO |
NO |
NO |
2.09 |
322.32 |
3 |
5 |
0 |
|
39 |
NO |
NO |
NO |
NO |
2.20 |
294.31 |
3 |
4 |
0 |
|
40 |
NO |
NO |
NO |
NO |
2.17 |
322.32 |
3 |
5 |
0 |
|
41 |
NO |
NO |
NO |
NO |
2.33 |
356.76 |
3 |
5 |
0 |
|
42 |
NO |
NO |
NO |
NO |
2.10 |
352.34 |
3 |
6 |
0 |
|
43 |
NO |
NO |
NO |
NO |
2.36 |
401.21 |
3 |
5 |
0 |
|
44 |
NO |
NO |
NO |
NO |
2.15 |
390.32 |
3 |
8 |
0 |
|
45 |
YES |
NO |
NO |
NO |
2.40 |
305.76 |
3 |
4 |
0 |
|
46 |
NO |
NO |
NO |
NO |
1.95 |
341.75 |
3 |
4 |
0 |
|
47 |
NO |
NO |
NO |
NO |
2.51 |
302.33 |
2 |
6 |
0 |
|
48 |
NO |
NO |
NO |
NO |
1.79 |
318.33 |
3 |
7 |
0 |
|
49 |
YES |
NO |
NO |
NO |
2.10 |
326.74 |
2 |
4 |
0 |
|
50 |
YES |
NO |
NO |
NO |
1.80 |
341.75 |
2 |
4 |
0 |
|
51 |
YES |
NO |
NO |
NO |
2.11 |
355.78 |
2 |
4 |
0 |
|
52 |
NO |
NO |
NO |
NO |
2.57 |
319.36 |
1 |
4 |
0 |
|
53 |
NO |
NO |
YES |
NO |
2.80 |
354.79 |
2 |
4 |
0 |
|
54 |
NO |
NO |
NO |
NO |
2.45 |
320.21 |
1 |
3 |
0 |
|
55 |
NO |
NO |
NO |
NO |
2.98 |
376.45 |
2 |
5 |
0 |
|
56 |
NO |
NO |
NO |
NO |
3.08 |
373.20 |
1 |
5 |
0 |
|
57 |
NO |
NO |
NO |
NO |
2.96 |
342.44 |
3 |
5 |
0 |
|
58 |
NO |
NO |
NO |
NO |
2.99 |
291.35 |
1 |
3 |
0 |
|
59 |
NO |
NO |
NO |
NO |
1.84 |
284.27 |
2 |
5 |
0 |
|
60 |
NO |
NO |
NO |
NO |
2.10 |
322.32 |
3 |
5 |
0 |
|
61 |
NO |
NO |
NO |
NO |
2.30 |
356.76 |
3 |
5 |
0 |
|
62 |
NO |
NO |
NO |
NO |
1.59 |
295.29 |
3 |
4 |
0 |
|
63 |
NO |
NO |
NO |
NO |
2.29 |
321.33 |
2 |
5 |
0 |
|
64 |
NO |
NO |
NO |
NO |
2.08 |
322.32 |
2 |
6 |
0 |
|
65 |
NO |
NO |
NO |
NO |
1.39 |
357.75 |
3 |
6 |
0 |
|
66 |
NO |
NO |
NO |
NO |
0.70 |
296.28 |
3 |
5 |
0 |
|
67 |
YES |
NO |
NO |
NO |
1.80 |
294.31 |
3 |
4 |
0 |
|
68 |
YES |
YES |
YES |
NO |
2.31 |
325.30 |
4 |
3 |
0 |
|
69 |
YES |
YES |
YES |
NO |
1.14 |
327.36 |
5 |
3 |
0 |
|
70 |
YES |
YES |
YES |
NO |
1.30 |
329.74 |
4 |
4 |
0 |
|
71 |
NO |
NO |
NO |
NO |
2.70 |
337.38 |
3 |
5 |
0 |
|
74 |
YES |
YES |
NO |
YES |
3.29 |
382.84 |
2 |
4 |
0 |
|
75 |
YES |
YES |
NO |
YES |
1.39 |
357.75 |
3 |
6 |
0 |
|
77 |
NO |
YES |
NO |
NO |
1.35 |
339.31 |
4 |
8 |
0 |
|
78 |
NO |
NO |
NO |
NO |
1.98 |
295.30 |
2 |
6 |
0 |
|
79 |
NO |
YES |
NO |
NO |
1.49 |
356.36 |
5 |
7 |
0 |
|
80 |
NO |
NO |
NO |
NO |
1.81 |
341.35 |
4 |
6 |
0 |
|
81 |
NO |
NO |
NO |
NO |
2.54 |
341.76 |
1 |
6 |
0 |
|
82 |
NO |
NO |
NO |
NO |
2.40 |
311.30 |
3 |
5 |
0 |
|
83 |
NO |
NO |
NO |
NO |
0.31 |
327.29 |
4 |
5 |
0 |
|
84 |
NO |
NO |
NO |
NO |
1.59 |
284.27 |
3 |
4 |
0 |
|
85 |
NO |
NO |
NO |
NO |
1.98 |
310.31 |
3 |
5 |
0 |
|
86 |
NO |
NO |
NO |
NO |
1.78 |
311.30 |
4 |
5 |
0 |
|
87 |
NO |
NO |
NO |
NO |
2.69 |
384.82 |
3 |
6 |
0 |
|
88 |
NO |
NO |
NO |
NO |
1.43 |
324.33 |
4 |
5 |
0 |
|
89 |
NO |
NO |
NO |
NO |
1.67 |
338.36 |
4 |
5 |
0 |
|
90 |
NO |
NO |
NO |
NO |
1.80 |
366.37 |
4 |
6 |
0 |
|
91 |
NO |
NO |
NO |
NO |
1.50 |
296.28 |
3 |
6 |
0 |
|
92 |
NO |
NO |
NO |
NO |
2.98 |
339.35 |
1 |
6 |
0 |
|
93 |
NO |
NO |
NO |
NO |
3.24 |
353.37 |
1 |
6 |
0 |
|
94 |
NO |
NO |
NO |
NO |
3.51 |
387.82 |
1 |
6 |
0 |
|
95 |
NO |
NO |
NO |
NO |
1.78 |
298.30 |
3 |
6 |
0 |
|
96 |
NO |
NO |
NO |
NO |
2.67 |
283.28 |
1 |
5 |
0 |
|
97 |
YES |
NO |
NO |
NO |
1.63 |
298.25 |
3 |
6 |
0 |
|
98 |
YES |
NO |
NO |
NO |
1.65 |
299.28 |
4 |
6 |
0 |
|
99 |
YES |
NO |
NO |
NO |
2.67 |
318.72 |
2 |
5 |
0 |
|
100 |
NO |
NO |
NO |
NO |
2.17 |
303.70 |
2 |
5 |
0 |
|
101 |
YES |
NO |
NO |
NO |
1.37 |
271.27 |
3 |
4 |
0 |
|
102 |
YES |
NO |
NO |
NO |
1.87 |
291.69 |
3 |
4 |
0 |
|
103 |
YES |
NO |
NO |
NO |
1.43 |
257.24 |
3 |
4 |
0 |
|
104 |
NO |
NO |
NO |
NO |
1.74 |
253.26 |
2 |
5 |
0 |
|
105 |
YES |
YES |
NO |
NO |
2.47 |
325.39 |
4 |
3 |
0 |
|
106 |
NO |
NO |
NO |
NO |
2.88 |
371.82 |
3 |
6 |
0 |
|
108 |
NO |
NO |
NO |
NO |
2.08 |
380.40 |
4 |
6 |
0 |
|
110 |
NO |
NO |
NO |
NO |
2.13 |
264.28 |
1 |
4 |
0 |
|
112 |
NO |
NO |
NO |
NO |
2.57 |
293.32 |
2 |
4 |
0 |
|
113 |
NO |
NO |
NO |
NO |
2.87 |
331.29 |
1 |
6 |
0 |
|
114 |
NO |
NO |
NO |
NO |
2.18 |
278.31 |
2 |
3 |
0 |
|
115 |
YES |
NO |
NO |
NO |
2.54 |
236.27 |
1 |
3 |
0 |
|
120 |
NO |
NO |
NO |
NO |
1.35 |
281.27 |
2 |
5 |
0 |
|
121 |
NO |
NO |
NO |
NO |
1.43 |
295.30 |
5 |
3 |
0 |
|
122 |
NO |
NO |
NO |
NO |
1.99 |
269.32 |
2 |
2 |
0 |
|
123 |
NO |
NO |
NO |
NO |
1.15 |
269.26 |
2 |
5 |
0 |
|
124 |
NO |
NO |
NO |
NO |
1.15 |
253.26 |
2 |
4 |
0 |
|
125 |
NO |
NO |
NO |
NO |
1.99 |
272.30 |
2 |
4 |
0 |
|
126 |
NO |
NO |
NO |
NO |
1.70 |
315.33 |
2 |
5 |
0 |
|
127 |
NO |
NO |
NO |
NO |
2.20 |
302.33 |
2 |
5 |
0 |
|
129 |
NO |
NO |
NO |
NO |
2.05 |
354.81 |
2 |
3 |
0 |
|
130 |
NO |
NO |
NO |
NO |
2.59 |
384.39 |
1 |
5 |
0 |
|
131 |
NO |
NO |
NO |
NO |
3.07 |
345.42 |
1 |
3 |
0 |
|
132 |
NO |
NO |
NO |
NO |
2.70 |
355.39 |
2 |
4 |
0 |
|
133 |
NO |
NO |
NO |
NO |
2.37 |
377.83 |
3 |
3 |
0 |
|
134 |
NO |
NO |
NO |
NO |
2.13 |
344.37 |
3 |
4 |
0 |
|
135 |
NO |
NO |
NO |
NO |
2.63 |
362.38 |
3 |
4 |
0 |
|
136 |
NO |
NO |
NO |
NO |
2.7 |
404.48 |
2 |
4 |
0 |
|
137 |
NO |
NO |
NO |
NO |
2.77 |
360.43 |
1 |
4 |
0 |
|
138 |
NO |
NO |
NO |
NO |
2.55 |
405.47 |
2 |
4 |
0 |
|
139 |
NO |
NO |
NO |
NO |
2.01 |
372.38 |
3 |
5 |
0 |
|
140 |
NO |
NO |
NO |
NO |
2.48 |
379.80 |
3 |
5 |
0 |
|
141 |
NO |
NO |
NO |
NO |
2.66 |
396.37 |
2 |
6 |
0 |
|
142 |
NO |
NO |
NO |
NO |
2.41 |
358.39 |
3 |
4 |
0 |
|
143 |
NO |
NO |
NO |
NO |
2.50 |
363.80 |
2 |
4 |
0 |
|
144 |
NO |
NO |
NO |
NO |
2.73 |
384.39 |
1 |
5 |
0 |
|
145 |
NO |
YES |
NO |
NO |
3.32 |
385.42 |
2 |
5 |
0 |
|
150 |
YES |
NO |
NO |
NO |
3.03 |
425.51 |
3 |
3 |
0 |
|
151 |
YES |
NO |
NO |
NO |
2.67 |
426.47 |
3 |
4 |
0 |
|
152 |
YES |
NO |
NO |
NO |
2.30 |
370.40 |
3 |
3 |
0 |
|
153 |
NO |
NO |
NO |
NO |
2.92 |
361.44 |
2 |
4 |
0 |
|
154 |
YES |
YES |
YES |
YES |
2.41 |
342.39 |
3 |
2 |
0 |
|
155 |
NO |
NO |
NO |
NO |
1.58 |
358.35 |
3 |
5 |
0 |
|
157 |
NO |
NO |
NO |
NO |
2.71 |
390.46 |
2 |
4 |
0 |
|
158 |
YES |
NO |
NO |
NO |
2.13 |
386.36 |
2 |
5 |
0 |
|
159 |
NO |
NO |
NO |
NO |
2.99 |
393.80 |
2 |
4 |
0 |
|
160 |
NO |
NO |
NO |
NO |
2.90 |
385.42 |
2 |
4 |
0 |
|
162 |
NO |
NO |
YES |
NO |
3.67 |
433.46 |
2 |
5 |
0 |
|
163 |
NO |
NO |
NO |
NO |
2.49 |
360.37 |
2 |
-5 |
0 |
|
164 |
NO |
NO |
NO |
NO |
2.66 |
341.36 |
2 |
4 |
0 |
|
166 |
NO |
NO |
NO |
NO |
2.44 |
412.40 |
1 |
6 |
0 |
|
170 |
NO |
NO |
NO |
NO |
3.43 |
373.83 |
1 |
3 |
0 |
|
171 |
NO |
NO |
NO |
NO |
2.94 |
381.84 |
2 |
3 |
0 |
|
172 |
NO |
NO |
NO |
NO |
2.61 |
370.36 |
1 |
5 |
0 |
|
173 |
NO |
NO |
NO |
NO |
2.65 |
467.50 |
2 |
6 |
0 |
|
174 |
NO |
NO |
NO |
NO |
2.95 |
356.38 |
2 |
4 |
0 |
MOLECULAR DOCKING
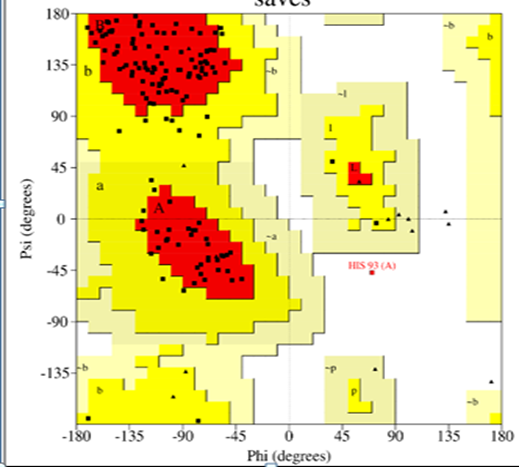
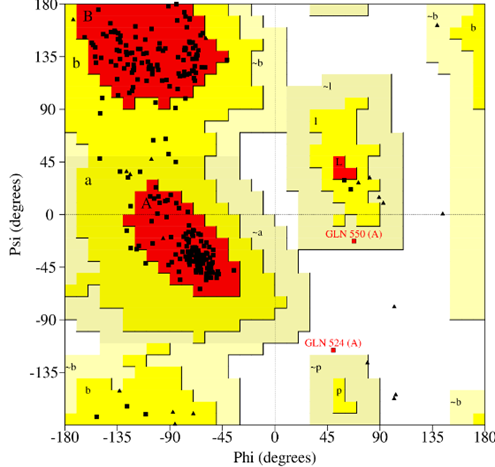
The structural integrity of DDR2 and ADAMTS-5 was confirmed through Ramachandran plot analysis (Fig. I), which showed that more than 85% for DDR2 and 90% for ADAMT5 of residues were located within the most favourable regions. Active site predictions for both targets were performed and are summarized in the results TABLE III. Ligands that demonstrated favorable drug-likeness and non-toxic profiles were selected for molecular docking against DDR2 and ADAMTS-5. Celecoxib was employed as the reference compound for comparative analysis of docking scores. The binding energies of the designed ligands ranged from –5.82 to –7.91 kcal/mol for DDR2, and from –6.63 to –9.56 kcal/mol for ADAMTS-5, indicating strong interactions with the target proteins.
|
7AZB |
2RJQ |
|
|
|
FIG NO.I: Ramachandran plot for DDR-2 &ADAMTS-5
|
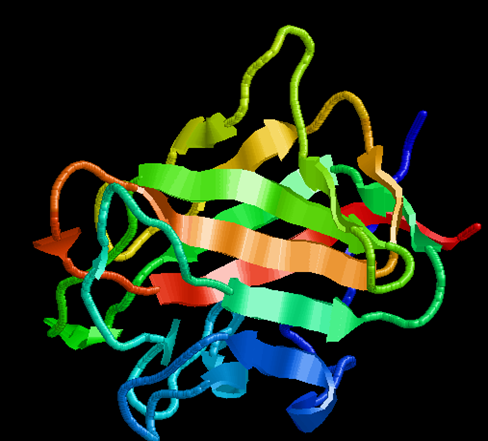
DDR-2(7AZB) |
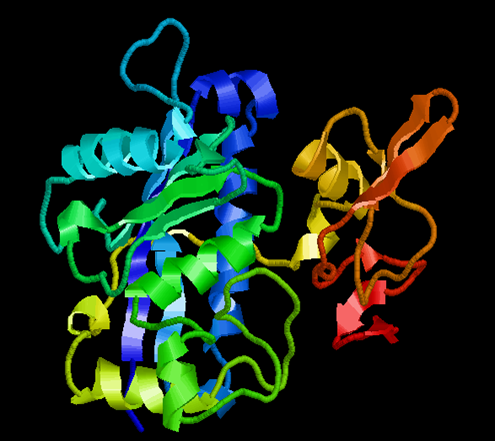
ADAMTS-5(2RJQ) |
|
|
|
FIG NO.II: 3D STRUCTURE OF PROTEIN BY RASMOL TOOL
Best predicted binding cavity for Molecular docking:
DDR-2: -20,0,16 (x,y,z)
ADAMTS-5: -39,-2, 20 (x,y,z)
TABLE NO. III: Binding scores of Ligands
|
S.NO |
LIGAND CODE |
DISCOIDIN DOMAIN RECEPTOR-2(7AZB) |
ADAMTS-5 (2RJQ) |
|
1) |
HMJ5 |
-6.98 |
-7.69 |
|
2) |
HMJ12 |
-6.28 |
-7.75 |
|
3) |
HMJ23 |
-7.64 |
-8.83 |
|
4) |
HMJ36 |
-6.23 |
-8.61 |
|
5) |
HMJ40 |
-5.82 |
-9.41 |
|
6) |
HMJ48 |
-7.51 |
-9.56 |
|
7) |
HMJ55 |
-6.40 |
-8.61 |
|
8) |
HMJ59 |
-7.17 |
-8.29 |
|
9) |
HMJ62 |
-7.21 |
-8.88 |
|
10) |
HMJ66 |
-7.91 |
-9.47 |
|
11) |
HMJ71 |
-6.51 |
-9.22 |
|
12) |
HMJ81 |
-7.85 |
-7.6 |
|
13) |
HMJ85 |
-7.55 |
-7.44 |
|
14) |
HMJ92 |
-7.37 |
-8.07 |
|
15) |
HMJ95 |
-6.54 |
-6.63 |
|
16) |
HMJ121 |
-7.31 |
-8.59 |
|
17) |
HMJ122 |
-5.90 |
-7.21 |
|
18) |
HMJ125 |
-6.10 |
-8.87 |
|
19) |
HMJ129 |
-7.63 |
-7.17 |
|
20) |
HMJ134 |
-7.53 |
-8.06 |
|
21) |
HMJ137 |
-6.89 |
-7.98 |
|
22) |
HMJ138 |
-7.72 |
-8.24 |
|
23) |
HMJ155 |
-7.46 |
-9.15 |
|
24) |
HMJ157 |
-7.13 |
-8.78 |
|
25) |
HMJ159 |
-5.94 |
-9.04 |
|
26) |
HMJ160 |
-6.03 |
-8.64 |
|
27) |
HMJ163 |
-7.18 |
-9.23 |
|
28) |
HMJ164 |
-7.03 |
-8.47 |
|
29) |
HMJ172 |
-7.49 |
-8.74 |
|
30) |
HMJ174 |
-7.33 |
-8.31 |
|
31) |
CELECOXIB |
-6.84 |
-7.23 |
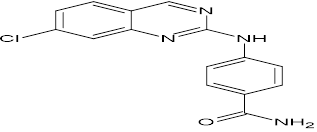
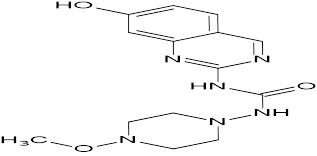
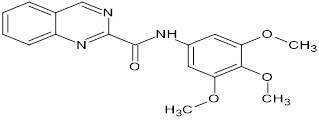
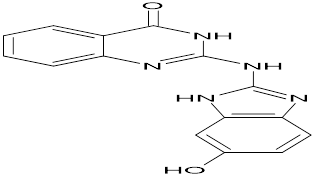
TABLE NO.IV: Chemical structure of Top-performing ligands based on docking Scores
|
LIGAND NO. |
STRUCTURE |
|
HMJ23 |
|
|
HMJ48 |
|
|
HMJ92 |
|
|
HMJ121 |
|
|
HMJ155 |
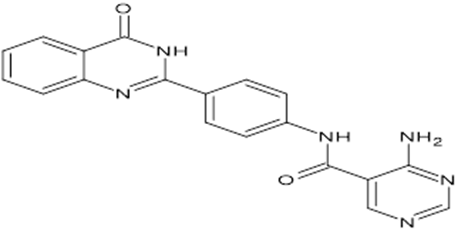
|
|
HMJ157 |
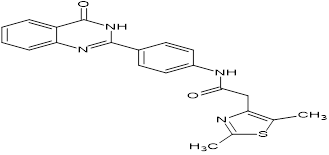
|
|
HMJ163 |
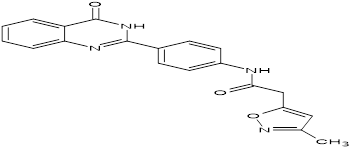
|
|
HMJ164 |
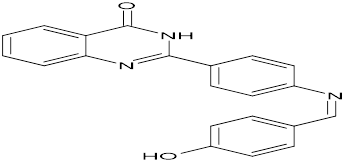
|
|
HMJ172
|
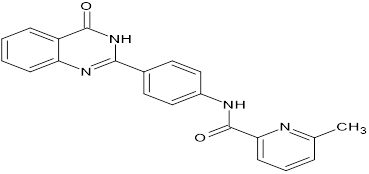
|
|
HMJ174 |
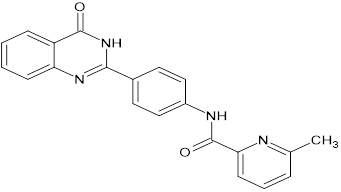
|
|
CELECOXIB |
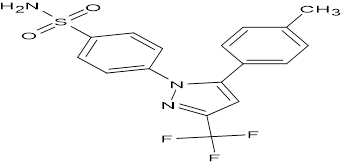
|
VISUALIZATION OF INTERACTIONS
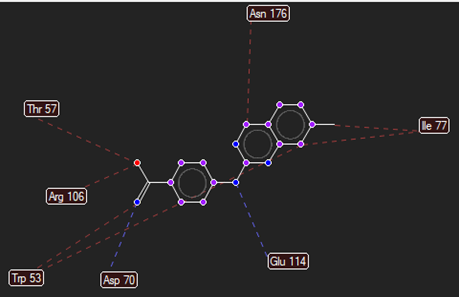
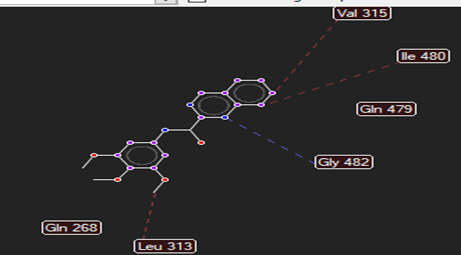
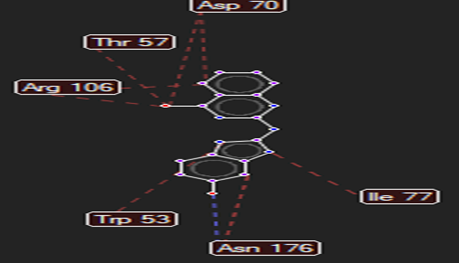
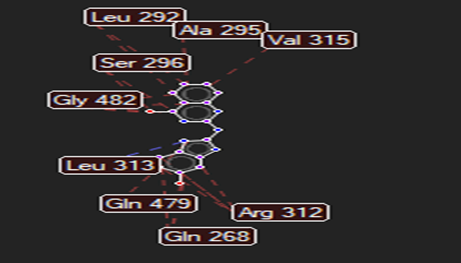
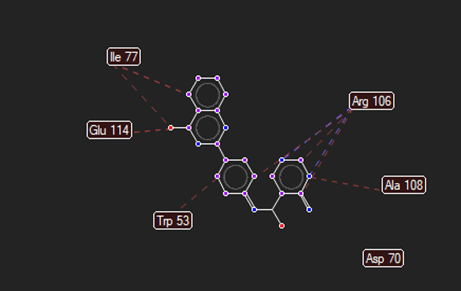
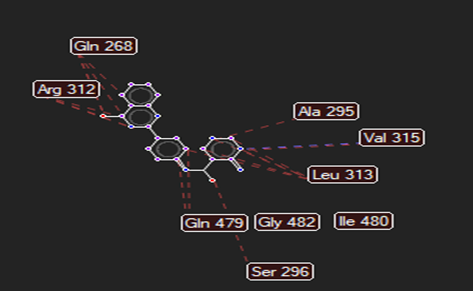
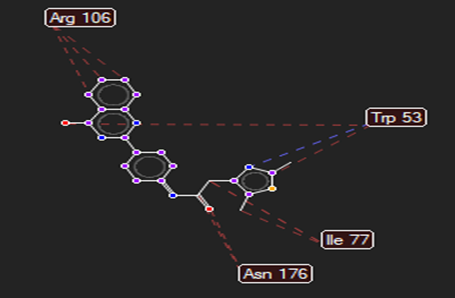
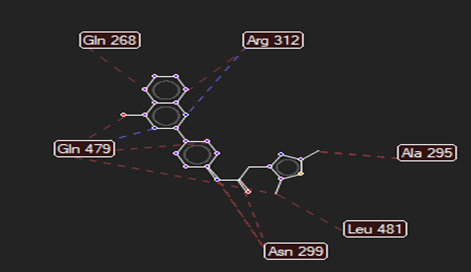
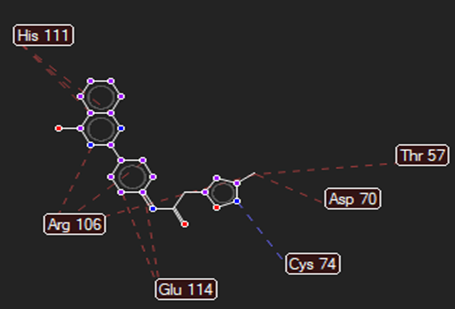
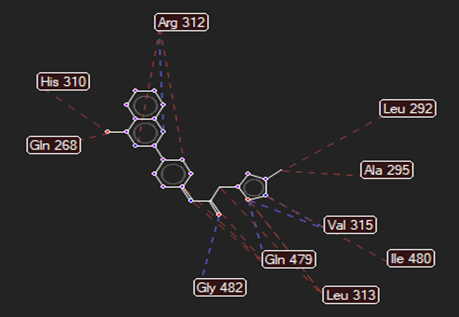
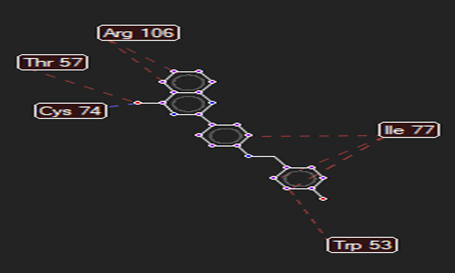
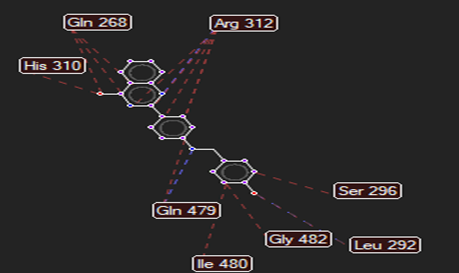
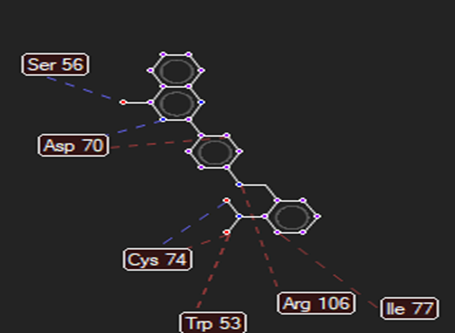
The ligand–receptor interactions were further analyzed and visualized, with the results summarized in TABLE V. Key interacting residues, types of interactions, and spatial relationships were studied to provide insights into the binding mechanisms of the selected ligands.
TABLE NO. V: Visualization of Ligand receptor interactions
|
LIG CODE |
DDR-2(7AZB)
|
ADAMTS-5(2RJQ) |
|
HMJ23 |
|
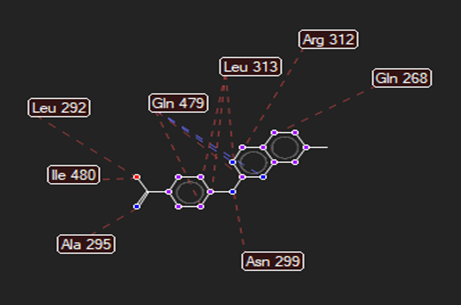
|
|
HMJ48 |
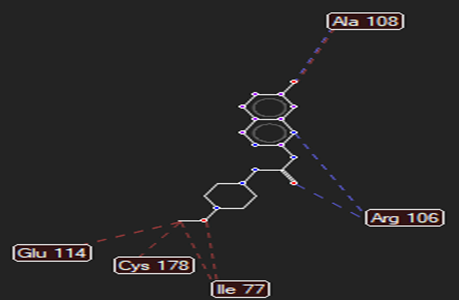
|
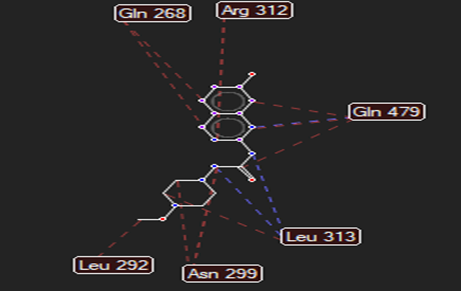
|
|
HMJ92 |
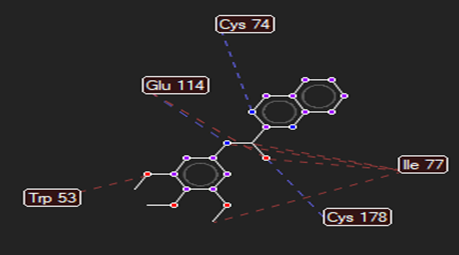
|
|
|
HMJ121 |
|
|
|
HMJ155 |
|
|
|
HMJ157 |
|
|
|
HMJ163 |
|
|
|
HMJ164 |
|
|
|
HMJ172 |
|
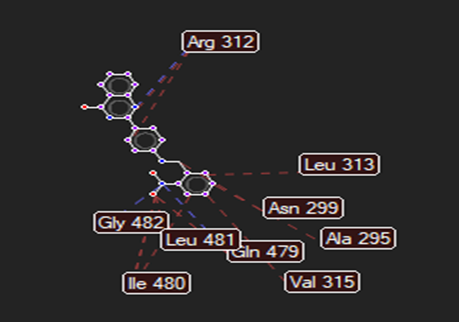
|
|
HMJ174 |
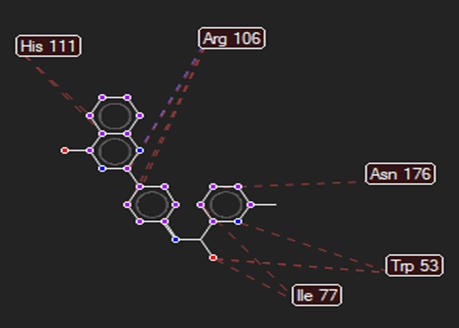
|
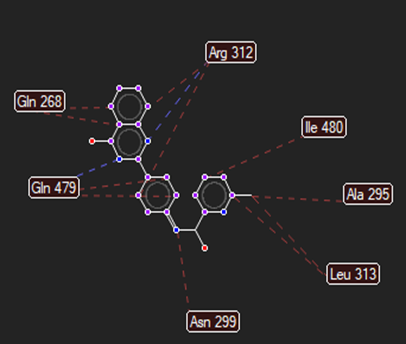
|
|
CELECOXIB |
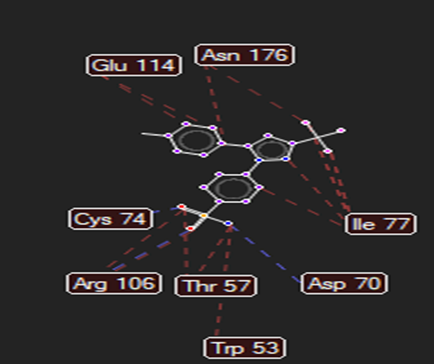
|
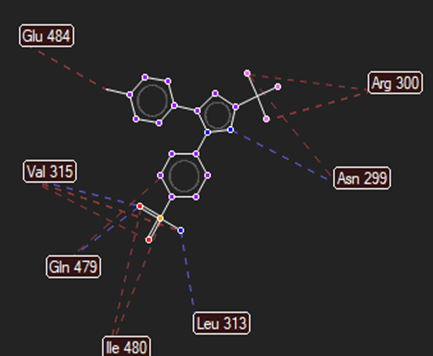
|
TABLE NO.V1: Ligand-receptor binding
|
LIGAND CODE |
HYDROGEN BONDING- DDR-2 |
HYDROGEN BONDING-ADAMTS-5 |
|
HMJ23 |
Asp70, Glu114 |
Gln 475 |
|
HMJ48 |
Arg106 |
Leu313 |
|
HMJ92 |
Cys78,Cys74,Gly114 |
Gly482 |
|
HMJ121 |
Asn176 |
Leu313 |
|
HMJ155 |
Arg106 |
Val315 |
|
HMJ157 |
Trp53 |
Gln479,Arg312 |
|
HMJ163 |
Cys74 |
Arg312,Gly482,Gln479, Ala295 |
|
HMJ164 |
Cys74 |
Arg312,Gln479 |
|
HMJ172 |
Ser56,Asp70,Cys74 |
Arg312,Gly482,Leu481 |
|
HMJ174 |
Arg106 |
Arg312,Gln479 |
|
CELECOXIB |
Arg106,Asp70 |
Val315,Leu35,Asn299 |
CONCLUSION
This study successfully demonstrates a rational, computer-aided drug discovery approach for identifying potential dual inhibitors targeting DDR2 and ADAMTS-5—two key enzymes implicated in the pathogenesis of osteoarthritis. A virtual library of 175 quinazolinone-based ligands was designed and screened using pharmacophore modeling, molecular docking, and ADMET predictions. The majority of compounds exhibited strong binding affinity, favorable drug-likeness, and low toxicity profiles. Notably, several ligands outperformed the reference drug Celecoxib in docking studies, suggesting superior therapeutic potential. These findings lay a promising foundation for the further development of disease-modifying osteoarthritis drugs (DMOADs) and support future in vitro and in vivo validation of the selected candidate.
ACKNOWLEGMENTS
We express our sincre thanks to the department of Pharmaceutical Chemistry, College of pharmacy, Madras Medical College(MMC), Chennai for providing necessary facilities for the Research work.
CONFLICTS OF INTEREST
The author declares there is no Conflict of Interest
REFERENCES




Priyadharshini R, Hanitha Mathanke J, Archana S, Guhan G, Dual Inhibition of DDR-2 and ADAMTS-5 by Novel 2-Substituted Quinazoline Derivatives: A Computational Approach for Disease-Modifying Osteoarthritis Drugs, Int. J. of Pharm. Sci., 2025, Vol 3, Issue 6, 5901-5916. https://doi.org/10.5281/zenodo.15774631
 10.5281/zenodo.15774631
10.5281/zenodo.15774631