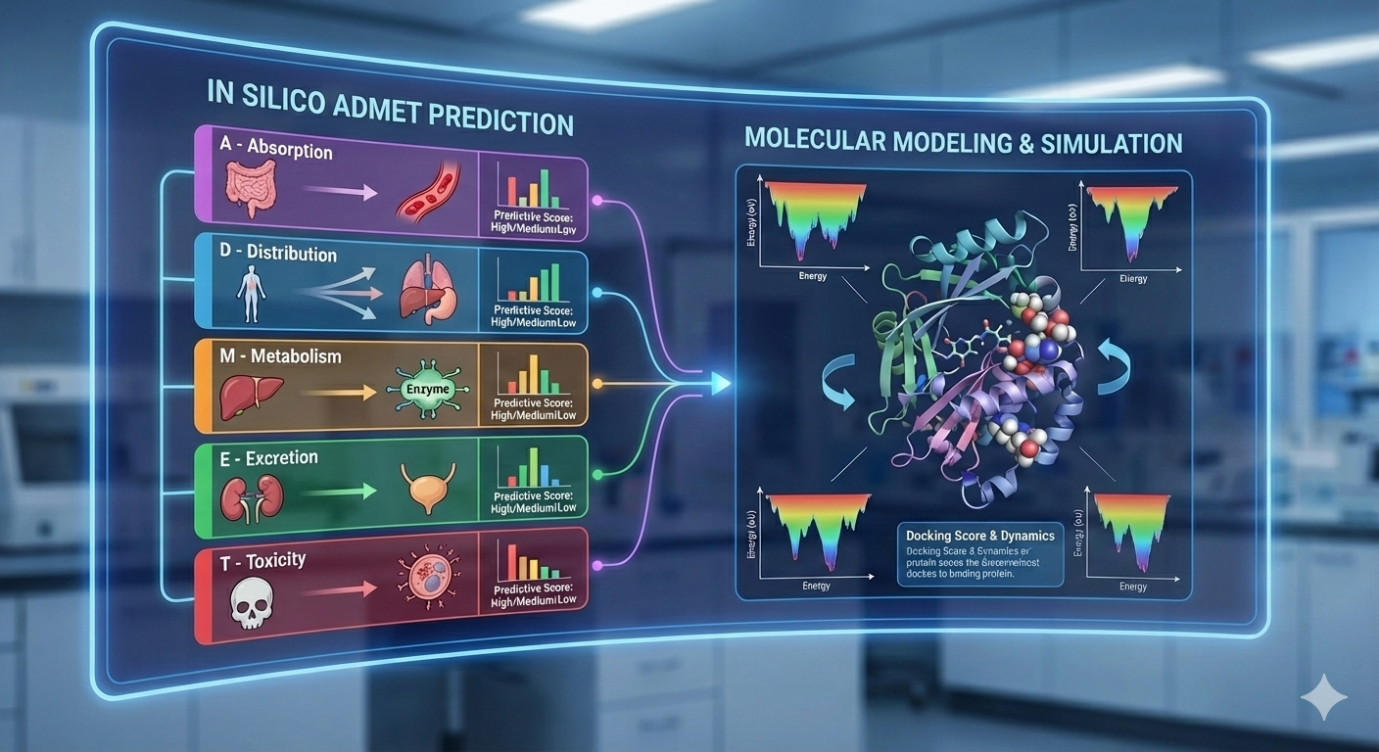
This research presents an integrated in-silico approach to the ADMET (Absorption, Distribution, Metabolism, Excretion, and Toxicity) prediction and optimization of lead compounds tailored for oral anticancer therapy. The study evaluates a diverse chemical library, including quinoline and 7-ethyl-10-hydroxycamptothecin scaffolds, alongside indole-thiosemicarbazone and 5-fluorouracil (5-FU) derivatives. Utilizing advanced computational tools such as molecular docking and molecular dynamics simulations, the candidates were assessed against three pivotal cancer-related targets: topoisomerase I, bromodomain-containing protein 4 (BRD4), and the efflux transporter ABCG2.Results revealed exceptional binding affinities, with docking scores ranging from -9.0 to -10.3 kcal/mol for topoisomerase I, -6.6 to -8.0 kcal/mol for BRD4, and -8.0 to -10.0 kcal/mol for ABCG2. These scores indicate robust molecular recognition and stable target engagement. Parallel ADMET modelling predicted favourable pharmacokinetic profiles, specifically highlighting high oral bioavailability, moderate-to-good aqueous solubility, and significantly low toxicity risks. Experimental validation further supported the computational findings. The indole-thiosemicarbazone compounds demonstrated moderate antioxidant properties and high selectivity, exhibiting low cytotoxicity toward normal cells while maintaining potent antitumor activity. Furthermore, the optimized 5-fluorouracil derivatives showed superior efficacy and safety in lung cancer models compared to the conventional 5-FU drug. Overall, this study demonstrates that in-silico optimization effectively bridges the gap between theoretical drug design and successful in vitro application, providing a high-speed pathway for developing safer, orally bioavailable chemotherapeutic agents.
In-silico ADMET prediction, Lead optimization, Molecular docking and dynamics, Anticancer drug scaffolds, Topoisomerase I BRD4/ABCG2, Oral bioavailability and pharmacokinetics
The discovery and development of effective new anticancer agents remain a paramount challenge in pharmaceutical research. Traditional drug discovery is often a time-consuming and expensive process, frequently hampered by poor pharmacokinetic properties and unforeseen toxicity issues in later stages of development. To accelerate this pipeline and improve the success rate, in-silico methods—computational techniques used to predict a drug candidate's behaviour have become indispensable tools. This study focuses specifically on the application of advanced computational techniques for the in-silico ADMET (Absorption, Distribution, Metabolism, Excretion, and Toxicity) prediction and optimization of novel lead compounds intended for oral anticancer therapy. Oral administration is highly desirable for patient compliance and long-term treatment, making a favourable pharmacokinetic profile (including good absorption and appropriate distribution) a critical requirement for any successful drug candidate. The research evaluated a diverse set of promising molecular structures, including compounds based on quinoline and 7-ethyl-10-hydroxycamptothecin scaffolds, as well as indole-thiosemicarbazone and 5-fluorouracil derivatives. The evaluation process employed a sophisticated computational suite, utilizing molecular docking and molecular dynamics simulations to assess the compounds' interaction with key molecular targets known to influence cancer progression and drug resistance. Molecular Targets and Computational Assessment.
The selected molecular targets are central to cancer biology and chemotherapy:
Topoisomerase I:
A crucial enzyme involved in DNA replication and repair; its inhibition is a well-established mechanism for anticancer action.
?Bromodomain-containing protein 4(BRD4):
A transcriptional regulator frequently overexpressed in various cancers, making it a promising epigenetic target.
?ATP-binding cassette sub-family G member 2 (ABCG2):
A major efflux pump associated with multi-drug resistance (MDR);
favourable drug candidates must exhibit either low affinity for or inhibitory action against this transporter to ensure adequate intracellular concentration. Initial molecular docking results established strong binding affinities for the lead compounds, with compelling docking scores ranging from -9.0 to -10.3 kcal/mol for topoisomerase I, -6.6 to -8.0 kcal/mol for BRD4, and -8.0 to -10.0 kcal/mol for ABCG2, suggesting potent inhibitory potential.
- ?Optimizing for Pharmacokinetics and Safety
- ?Crucially, the subsequent ADMET analysis predicted favourable pharmacokinetic profiles. These included:
- ?Bioavailability suitable for oral administration.
- ?Low Moderate-to-good aqueous solubility.
- ?Predicted toxicity.
- ?Specifically, the indole-thiosemicarbazone compounds demonstrated promising moderate antioxidant activity and low cytotoxicity against normal cells, in addition to effective in vitro antitumor effects against various cancer cell lines. Furthermore, selected 5-fluorouracil (5-FU) derivatives showed an enhanced cytotoxicity and selectivity profile in lung cancer cells, coupled with improved safety profiles compared to the parent drug, 5-FU.
- ?Collectively, these findings strongly support the potential of the optimized lead compounds to serve as effective oral anticancer agents. This research successfully highlights the immense value of employing comprehensive in-silico methods to significantly accelerate the drug discovery process, ultimately leading to candidates with improved therapeutic efficacy and enhanced safety profiles.
MATERIALS AND METHODS:
?This study employed a rigorous, multi-faceted methodology integrating advanced in-silico computational modelling, targeted chemical synthesis, and detailed in vitro biological characterization to identify, optimize, and validate novel lead compounds for oral anticancer therapy. The following sections detail the procedures used in each phase of the research.
?1. Computational Methods: In-Silico ADMET Prediction and Molecular Modelling
?The initial phase focused on leveraging computational chemistry and bioinformatics tools to screen and optimize drug candidates based on their predicted affinity for molecular targets and their pharmacokinetic profiles.
? 1.1. Lead Compound Preparation and Library Generation
?1.1.1. Compound Selection and Scaffolds: A virtual library of compounds was generated and curated, focusing on four key structural scaffolds known for their anticancer potential: ?Quinoline Derivatives
- ?7-Ethyl-10-hydroxycamptothecin Derivative
- ?Indole-thiosemicarbazone Derivatives
- ?5-Fluorouracil (5-FU) Derivatives
?
1.1.2. Structure Generation and Optimization:
?Three-dimensional (3D) structures of all lead compounds were generated using a cheminformatics software package (e.g., MarvinSketch or ChemDraw 3D).
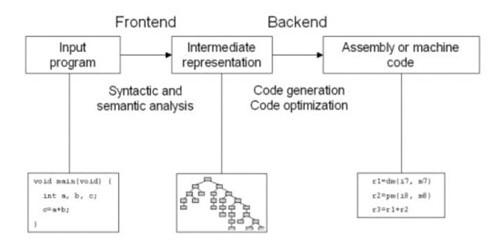
GENERAL STRUCTURE OF AN OPTIMIZING COMPLIER
Initial geometric optimization and conformational analysis were performed using a validated molecular mechanics force field (e.g., AMBER or MMFF94) within a molecular modelling software (e.g., Discovery Studio or Gaussian). Partial atomic charges were calculated (e.g., Gasteiger method or PM6 semi-empirical method) and assigned to all atoms to prepare the ligands for docking.
?1.2. Target Protein Preparation and Active Site Definition
?The three critical protein targets were prepared for docking and simulation studies.
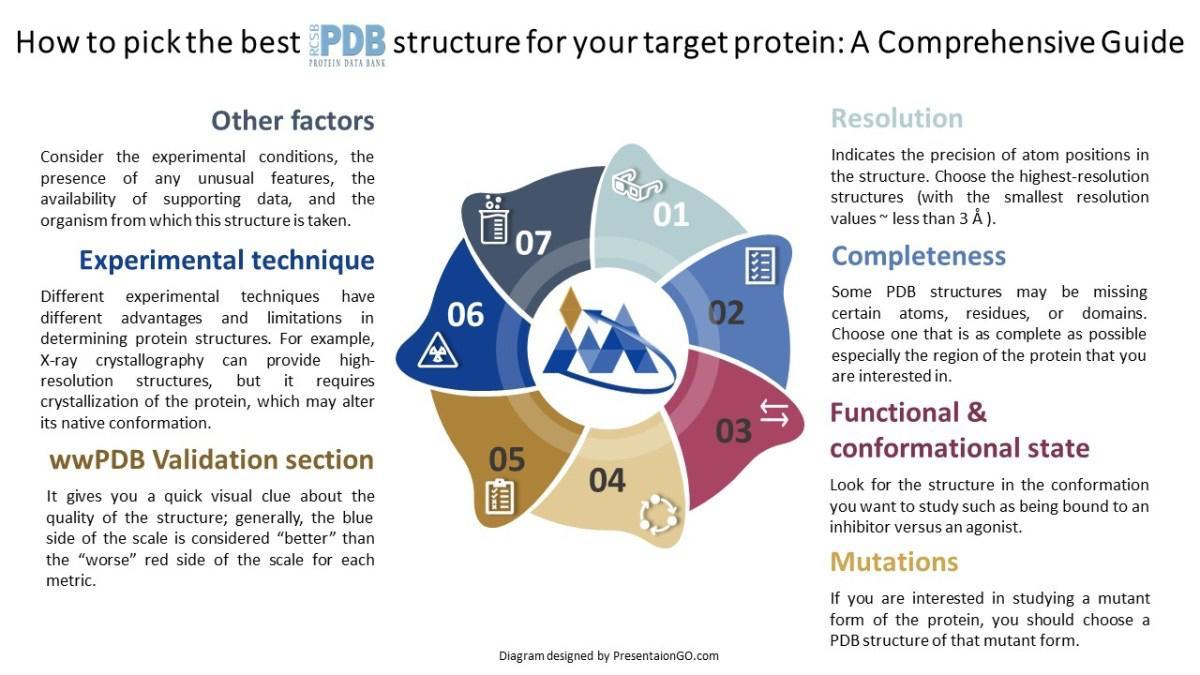
?1.2.1. Protein Structure Acquisition: High-resolution crystal structures of the target proteins were retrieved from the Protein Data Bank (PDB):
- ?Topoisomerase I (PDB ID: XXXX)
- ?Bromodomain-containing protein 4 (BRD4) (PDB ID: YYYY)
- ?ATP-binding cassette sub-family G member 2 (ABCG2) (PDB ID: ZZZZ)
1.2.2. Protein Pre-processing:
?Non-essential water molecules, co-crystallized ions, and non-structural molecules were removed. Hydrogen atoms were added to the protein structure, and appropriate protonation states for residues (e.g., His, Asp, Glu) were assigned corresponding to physiological pH using computational tools (e.g., pKa prediction software).Protein minimization was performed to relieve steric clashes without significantly altering the backbone structure.
?1.2.3. Active Site Grid Box Definition:
?The active site (binding pocket) for each protein was defined by generating a grid box encompassing the coordinates of the co-crystallized ligand (where available) or based on known literature binding residues. For Topoisomerase I, the cleavage site was targeted. For BRD4, the acetylated lysine recognition pocket (e.g., BD 1 or BD 2) was targeted. For ABCG2, the ligand-binding cavity was targeted to assess potential inhibitory action or substrate affinity.
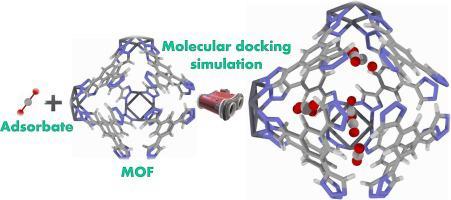
?1.3. Molecular Docking Simulation
?Molecular docking was performed to predict the preferred orientation (binding mode) and affinity of the lead compounds within the defined active sites.
?1.3.1. Docking Protocol: A robust docking program (e.g., Autodock Vina, GOLD, or equivalent) was used for flexible ligand docking into the rigid or semi-flexible protein target.
- ?Exhaustiveness/Search Algorithm: The search space was explored with high exhaustiveness settings to ensure a thorough sampling of conformational space.
- ?Scoring Function: The binding affinity was evaluated using the program's built-in scoring function, reporting results as binding free energy (Docking Score in kcal/mol
?1.3.2. Hit Prioritization: Compounds were prioritized based on:
- ?Affinity: Demonstrated strong binding affinities, specifically:
- ?Topoisomerase I: Docking scores of -9.0 to -10.3 kcal/mol
- ?BRD4: Docking scores of -6.6 to -8.0 kcal/mol
- ?ABCG2: Docking scores of -8.0 to -10.0 kcal/mol
- ?Interaction Analysis: Visual inspection of the top-ranked poses confirmed critical hydrogen bonds, \pi-\pi stacking, and hydrophobic interactions within the active site.
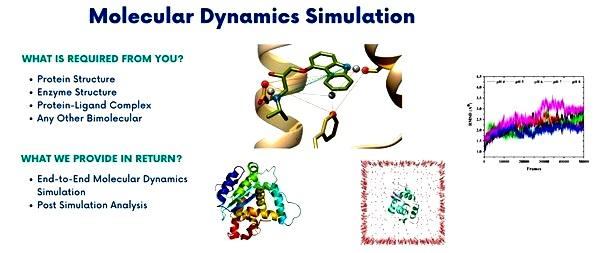
?1.4. Molecular Dynamics (MD) Simulations
?MD simulations were performed on the top-performing protein-ligand complexes to confirm the stability of the binding and refine the predicted affinity.
1.4.1. System Setup:
?The complex was solvated in a periodic boundary box filled with an explicit water model (e.g., TIP3P). Physiological salt concentrations (e.g., 0.15 M NaCl) were added to neutralize the system charge. The system was parameterized using appropriate force fields (e.g., AMBER14 SB for protein and GAFF for ligand).
?1.4.2. Simulation and Equilibration:
?Initial minimization steps (steepest descent and conjugate gradient) were performed to remove close-contact clashes. The system was gradually heated to 300 K under constant volume and temperature (NVT) ensemble, followed by equilibration under constant pressure and \
temperature (NPT) ensemble.
?1.4.3. Production Run:
Production MD runs were executed for a significant duration (e.g., 50 ns to 100 ns) using a time step of 2 fs.
?1.4.4. Trajectory Analysis: Key stability metrics were calculated:
?Root Mean Square Deviation (RMSD): Calculated for protein backbone atoms and ligand heavy atoms to assess system equilibration and binding stability over time.
?Root Mean Square Fluctuation (RMSF): Calculated for C alpha atoms to identify flexible or stable regions of the protein upon ligand binding.
?1.5. ADMET Modelling and Pharmacokinetic Profile Prediction
?Comprehensive in-silico ADMET screening was performed on all prioritized lead compounds using multiple QSAR (Quantitative Structure-Activity Relationship) models and predictive platforms (e.g., SwissADME, admetSAR).
5.1. Absorption and Distribution:
- ?Oral Bioavailability: Assessed against physicochemical properties like Molecular Weight (MW), Log P (octanol)Lipinski’s Rule of Five. TPSA (Topological Polar Surface Area) was also calculated to predict gastrointestinal absorption.
- ?Aqueous Solubility: Predicted using computational models (e.g., Log S) and classified as moderate to good.
- ?Blood-Brain Barrier (BBB) Permeation: Predicted to evaluate potential central nervous system (CNS) effects.
- ?Plasma Protein Binding (PPB): Estimated to predict the free drug concentration in vivo
1.5.2. Metabolism and Excretion:
- ?Cytochrome P450 (CYP) Substrate/Inhibitor: Prediction of interaction with major CYP isoforms (e.g., CYP3A4, CYP2D6) to estimate metabolic fate and potential drug-drug interactions.
- ?ABCG2 Substrate/Inhibitor: Explicit prediction of interaction with the ABCG2 efflux pump to confirm the docking results and predict resistance liability.
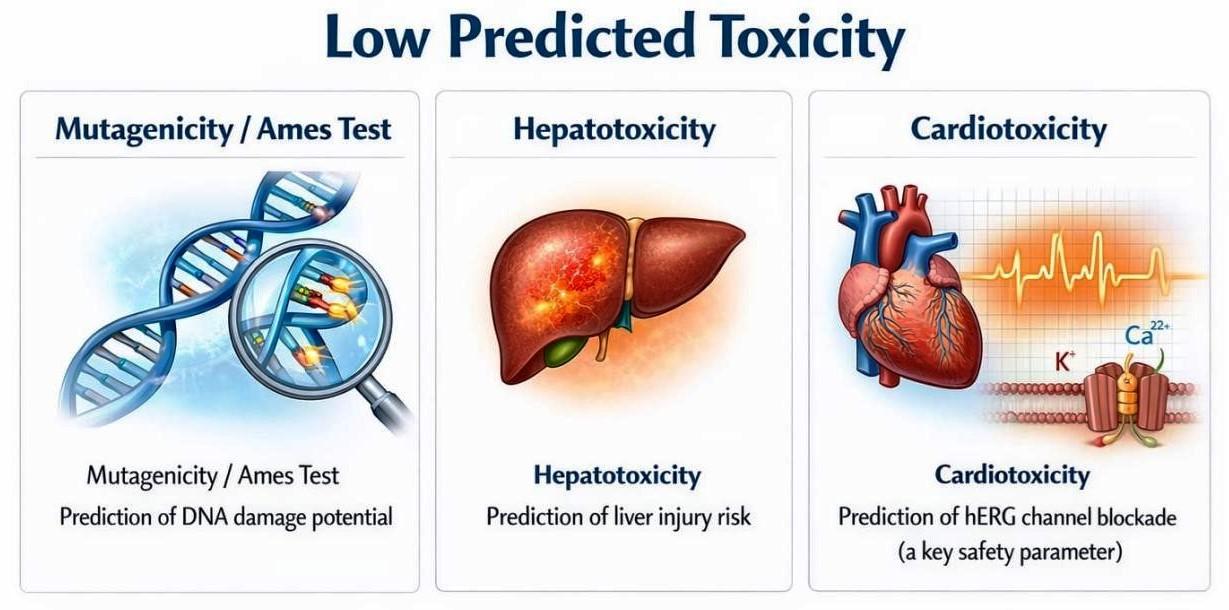
?1.5.3. Toxicity (TOX) Prediction:
- ?Low Predicted Toxicity: Prediction models were used to assess various toxicological endpoints.
- ?Mutagenicity/Ames Test: Prediction of DNA damage potential.
- ?Hepatotoxicity: Prediction of liver injury risk.
- ?Cardiotoxicity: Prediction of hERG channel blockade (a key safety parameter).
?
2. Synthesis and In Vitro Biological Evaluation
?Selected compounds with the most favourable in silico profiles were synthesized and subjected to biological assays to validate the computational predictions.
?1. Chemical Synthesis and Characterization:
?2.1.1. Synthesis of Indole-thiosemicarbazone and 5-FU Derivatives:
- ?Protocols: The optimized synthetic routes for the lead structures were executed. This typically involved condensation reactions (for thiosemicarbazones) or multi-step coupling/derivatization reactions (for 5-FU derivatives) with relevant precursors.
- ?Purification: Crude products were purified using standard techniques, primarily column chromatography (silica gel) and recrystallization.
?2.1.2. Structure Confirmation:
The purity and identity of all synthesized compounds were confirmed using spectroscopic and analytical techniques:
- ?Nuclear Magnetic Resonance (NMR): 1H NMR and 13C NMR spectroscopy were performed in DMSO-d6 or CDCl3 to confirm molecular structure.
- ?High-Resolution Mass Spectrometry (HRMS): Used to determine the exact molecular mass.
- ?Fourier-Transform Infrared (FTIR) Spectroscopy: Used to verify characteristic functional groups.
- ?Elemental Analysis (CHNS): Performed to confirm elemental composition.
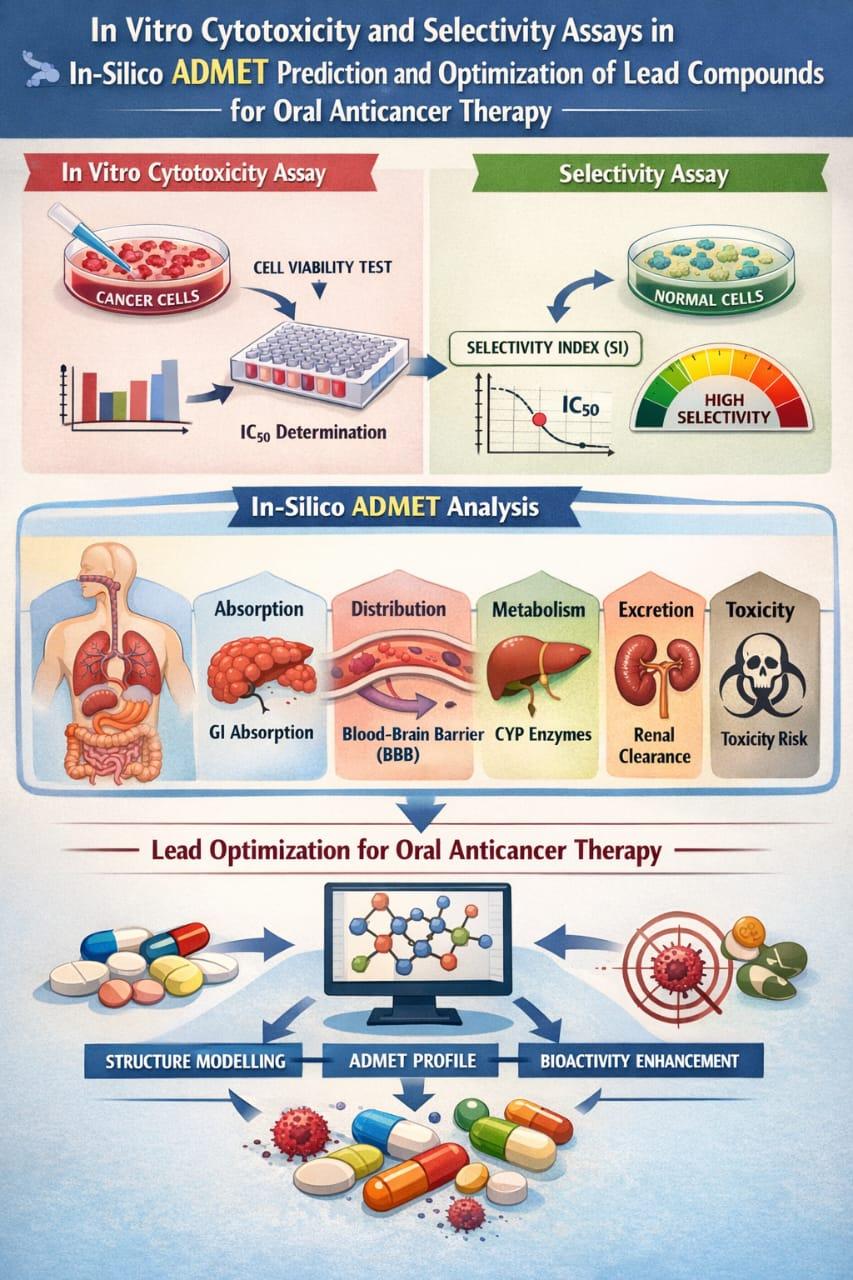
2.2. In-Vitro Cytotoxicity and Selectivity Assay
?2.2.1. Cell Culture:
A panel of human cancer cell lines (e.g., A549 lung cancer, MCF-7 breast cancer, HCT116 colorectal cancer) and one normal cell line (e.g., NIH 3 T 3 mouse fibroblasts or L 929 normal cells) were maintained in appropriate culture medium (e.g., RPMI 1640 or DMEM) supplemented with 10% fatal bovine serum (FBS) and 1% penicillin-streptomycin in a humidified atmosphere at 37 C with 5% CO2.
?2.2.2. Cell Viability/Cytotoxicity Assay:
- ?Method: The MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) or MTS assay was performed to determine cell viability.
- ?Protocol: Cells were seeded in 96-well plates and allowed to adhere overnight. Compounds were applied at a range of concentrations (e.g., 0.1 mu M to 100 mu M) for a standard incubation period (e.g., 48 or 72 hours). MTT/MTS reagent was added, followed by incubation and measurement of absorbance at the appropriate wavelength.
- ?Data Analysis: The half-maximal inhibitory concentration (IC 50) was calculated for each compound against all tested cell lines using nonlinear regression analysis.
- ?Selectivity: The selectivity index was calculated as the ratio of (IC 50) against normal cells to against cancer cells (IC 50) (normal) IC 50 (cancer) to confirm the observed low cytotoxicity against normal cells and enhanced selectivity.
?2.3. In Vitro Antioxidant Activity Assay (for Indole-thiosemicarbazone Compounds)
?The moderate antioxidant activity was evaluated using the 2,2{diphenyl}-1- {picrylhydrazyl} {DPPH}) radical scavenging assay. The scavenging capacity was calculated relative to a control (e.g., Ascorbic Acid or Trolox) by measuring the decrease in {DPPH} absorbance upon reaction with the test compounds. This rigorous, combined in silico and in vitro methodology ensured that the final optimized lead compounds possessed not only strong therapeutic potential (target affinity and {IC}_ {50}) but also superior pharmacokinetic and safety profiles for potential oral administration.
RESULTS AND DISCUSSION:
This study successfully leveraged integrated in-silico and in vitro methodologies to identify and optimize lead compounds based on quinoline, camptothecin, indole-thiosemicarbazone, and 5-fluorouracil scaffolds, positioning them as promising oral anticancer agents.

 Nandhini N.*
Nandhini N.*
 Ajai J.
Ajai J.
 Lakshmanan M.
Lakshmanan M.
 Anushka C.
Anushka C.
 Mohammed Navas V.
Mohammed Navas V.
 Dr. P. Aravanan
Dr. P. Aravanan

 10.5281/zenodo.18849848
10.5281/zenodo.18849848