
We use cookies to ensure our website works properly and to personalise your experience. Cookies policy

We use cookies to ensure our website works properly and to personalise your experience. Cookies policy
Head of Solutions Architecture, Regeneron.
The Life Sciences industry has started to transform its enterprise data architecture through three stages which began with data lakes followed by data products and now lakehouse architecture. The review examines multiple developments which focus on moving from single storage systems to specialized domain-based data solutions throughout 2019-2025. The authors considered technical foundations, challenges, and potential for the models to create change using emergent findings from 15 peer-reviewed studies, contemporaneous research findings, and industry reports. Life Sciences organizations are going all in on lakehouse architectures that combine the best of data lakes and data warehouses by textbook definitions but operate at nearly double the efficiencies of their legacy systems. The global market of the Life Sciences data mesh will reach $8.98 billion by 2033 while growing at an annual growth rate of 19.4%. In 2025, 70% of new analytics projects will leverage lakehouse capabilities. Moreover, 22% employ domain-specific AI tools. However, key challenges will still be addressed such as interoperability, workforce gaps in FHIR standards, and integration with legacy systems. Life Sciences organizations need to modernize, govern, upskill, and adopt emerging interoperability standards if they are to become productive for precision medicine and real-time insights.
The Life Sciences industry generates a huge amount of varied data from sources like electronic health records, imaging systems, lab information systems, wearable technology, and genomic sequencing platforms. Keeping up with the rapid increase in this data has become one of the biggest challenges for Life Sciences organizations today. Recent research suggests that global Life Sciences data could grow at an incredible rate of 36% annually by 2025, which is much faster than Moore's Law predicts [1] Yet, frustratingly, less than 3% of this valuable data is being effectively used for analysis and decision-making right now. That only means there are a lot of missed opportunities that we need to capture, so we will have better research outcomes and delivery of data-driven innovation across Life Sciences domains. Now, in health information management, there are two kinds of options being pursued: centralized data storage systems or data lakes. The query performance of data warehouses remains strong, but their adaptability to different data formats and fast-changing data types remains limited. Data lakes operate as expandable storage systems which accept unprocessed data in any format.
The main issue with data lakes stems from poor data quality and governance problems which make it difficult to extract valuable insights and results in what experts call 'data swamps' [2]. The existing separation between data storage systems continues to create operational problems which restrict Life Sciences organizations from achieving full data resource utilization. The Life Sciences industry faces ongoing data management challenges which data lakehouses and data mesh frameworks aim to solve through their combination of warehouse and lake advantages and their domain-based decentralized data governance models. The Life Sciences analytics market continues to expand at a 21.1% annual rate until 2030 because organizations need appropriate data infrastructure to support artificial intelligence applications and real-time clinical decision support artificial intelligence applications for drug discovery, clinical trial optimization, translational research, and biomarker analytics initiatives [3]. Organizations that need to understand these new architectural designs because they will determine how Life Sciences organizations can use data strategically in their digital transformation journey.
Life Sciences organizations experience an extreme data management emergency because their systems operate independently from each other while using different data weaknesses in Life Sciences systems. The COVID-19 data reporting across 36 states became inconsistent during mid- formats and lacking proper infrastructure for complex analytics and artificial intelligence operations. The COVID-19 pandemic exposed fundamental2021 while vaccine recipient race and ethnicity data remained unreported by states and jurisdictions at less than 50% which created major doubts about vaccine distribution fairness [4]. The public health response and health equity efforts became limited because of insufficient data infrastructure. The current Life Sciences data systems encounter three major operational challenges. The main problem with Life Sciences data systems stems from different technological systems and data formats which fail to meet interoperability standards despite regulatory requirements [5]. The current centralized data systems face several challenges in scaling up their operations, as Life Sciences data sources grow rapidly in numbers and diversity, hindering real-time analytics for clinical decision support [6].
Added to these technical issues in Life Sciences research environments are increasingly complex challenges related to governance and security, particularly with respect to compliance with Health Insurance Portability and Accountability Act (HIPAA) standards for the protection of research information. The estimated five- and ten-year costs of these infrastructure problems are, respectively, 7.84 billion dollars and 36.7 billion dollars, although neither estimate includes the full cost of an enterprise clinical data system. The true cost of inaction is most likely far higher. Inaccurate data, inefficient analysis pipelines, delayed research insights, and fewer opportunities to advance precision medicine research can all be caused by inadequate data infrastructure. For Life Sciences enterprises wishing to raise the quality of research and data-driven decision-making and keep competitive in the information-intensive environment of today, updating data systems is not an option; it's a necessity. To improve clinical trial outcomes, this modernization gives equal emphasis to improved economic management and technology.
Objectives
The research is ongoing, and it investigates the evolution of health data management from the outdated notion of data lakes to modern data products and lakehouse systems. This research has been focused on the analysis of technical features and designs related to data lakes, warehouses, and lakehouses; studying current trends in the implementation of these respective data architectures within Life Sciences institutions; and establishing the challenges these organizations have found during the deployment of advanced data systems. Finally, the analysis performed below outlines the trends and challenges of global Life Sciences data management. Fourth, to determine the place of interoperability standards, especially Fast Life Sciences Interoperability Resources, in facilitating efficient data sharing and integration across contemporary data architectures. Fifth, to examine artificial intelligence and machine learning capabilities that are being integrated into contemporary Life Sciences data platforms and their impacts on clinical and operational outcomes. Sixth, this report synthesizes best practices, lessons learned, and strategic recommendations for Life Sciences organizations undertaking data architecture modernization initiatives. Finally, directions for future research and development in Life Sciences data architecture propose information gaps in the current state of knowledge and new emerging opportunities for innovation in this rapidly changing field. This shows the stepwise screening process used in this systematic review. Of the 127 initially identified records, tracking citations and including workshops increased this number to 168 papers. Title/abstract screening resulted in a list of 58 papers, which was further reduced to 28 after examination of introductions and methods, and finally, 15 studies were included for full analysis.
Approach
This systematic review uses a comprehensive literature analysis method to study the state of evolution of Life Sciences data architecture. Accordingly, the review adopted a structured approach by comprising two important phases: First, a comprehensive database search in academic repositories, industry publications, and Life Sciences technology reports for sources published within 2019-2025 was performed. The search terms comprise various combinations of data lakes, data products, data lakehouse, data mesh, Life Sciences data architecture, enterprise data management, FHIR interoperability, and Life Sciences analytics. The selection process focused on peer-reviewed research articles and industry reports from reputable Life Sciences technology organizations and market analysis documents from established firms and empirical case studies that document actual implementation scenarios. The resources focus on Life Sciences research applications instead of enterprise data systems and they need to provide concrete data instead of abstract concepts and they must present new findings instead of summarizing existing information.
The different analytical phases required researchers to extract particular details about technical architecture and implementation approaches and numerical data and performance results and study outcomes and market patterns and user adoption rates. The research combined data from various sources to identify common patterns and emerging patterns and conflicting viewpoints between different studies. The review also considered a number of architecture philosophies and established their advantages by respective examples. Alongside operational and strategic aspects, the technical aspects provide a comprehensive understanding of the current state of health data architecture and its probable paths of future development.
Here is a chronological summary in table 1 of 15 papers included in this systematic review in order to obtain a synopsis of research evolution alongside key contributions towards the field.
Table 1: Chronological Summary of Reviewed Papers
|
Year |
Paper Title |
Key Findings |
Ref |
|
2019 |
Data Lakehouse Life Sciences Operations |
Life Sciences data growth 36% annually; 70% adoption prediction by 2025 |
[1] |
|
2020 |
Data Lakehouse Architecture Benefits |
Combines flexibility with ACID compliance; real-time analytics capabilities |
[2] |
|
2020 |
Life Sciences Data Platform Evolution |
Life Sciences analytics market CAGR 21.1% through 2030 |
[3] |
|
2021 |
National Health Data Enterprise Architecture |
USD 7.84-36.7B needed for infrastructure modernization |
[4] |
|
2022 |
Life Sciences Integration and Interoperability |
78% of providers using FHIR report faster coordination |
[5] |
|
2022 |
Data Lakehouse Implementation Examples |
Montefiore PALM semantic data lake for AI at scale |
[6] |
|
2023 |
Life Sciences Data Lake Benchmarking |
Architecture design and performance evaluation frameworks |
[7] |
|
2023 |
Lakehouse Platform Performance |
2x speed improvement; daily data refresh in minutes |
[8] |
|
2024 |
AI Adoption in Life Sciences |
22% implemented AI tools; USD 1.4B spending in 2024 |
[9] |
|
2024 |
Data Mesh Market Trends |
USD 1.3B in 2023; 17% CAGR to 2030 |
[10] |
|
2024 |
FHIR Data Models Systematic Review |
Pipeline-based and static model implementations analyzed |
[11] |
|
2024 |
Microsoft Life Sciences Data Solutions |
Bronze-silver-gold medallion lakehouse architecture |
[12] |
|
2025 |
Life Sciences Data Mesh Market |
USD 1.82B in 2024; 19.4% CAGR to USD 8.98B by 2033 |
[13] |
|
2025 |
FHIR Implementation Barriers |
Limited workforce knowledge; incomplete EHR vendor support |
[14] |
|
2025 |
MLOps in Life Sciences |
MLOps architecture with automation and reproducibility |
[15] |
Research Questions
This review is directed by seven main research questions that shape the examination and analysis.
RQ1: What are the core architectural distinctions among traditional data lakes, data warehouses, contemporary data lakehouses, and new data mesh patterns, and how do these variations appear in Life Sciences applications?
RQ2: What is the present status of modern data architecture adoption in Life Sciences organizations worldwide, and what elements are promoting or hindering this adoption?
RQ3: What are the main technical, organizational, and regulatory hurdles that Life Sciences organizations face when adopting data lakehouse or data mesh architecture?
RQ4: In what ways do Fast Life Sciences Interoperability Resources and various interoperability standards converge with contemporary data architectures, and what challenges hinder the attainment of semantic interoperability on a large scale?
RQ5: The current Life Sciences data systems implement artificial intelligence and machine learning through specific operational methods which architectural decisions either support or limit artificial intelligence functionality.
RQ6: Life Sciences organizations that implemented modern data architectures achieved measurable improvements in operational performance and clinical results and operational efficiency.
RQ7: Life Sciences organizations can enhance their data infrastructure through specific best practices and design patterns and strategic approaches.
Architectural Evolution and Comparison
The architectural transformation has occurred because of technological developments and organizational requirements. The schema-on-write approach in traditional data warehousing enables fast query execution and delivers high-quality results. The system faces limitations in handling different data types because its extract-transform-load process takes a long time which reduces its agility. The flexibility of data lakes is restricted because they store unprocessed diverse data in large quantities, but they create data swamps that make information retrieval and reuse challenging [9]. The lakehouse architecture helps deal with the individual weaknesses of warehouses and lakes by combining their advantages. The lakehouse architecture provides warehouse-grade ACID transactions and schema enforcement and governance capabilities alongside lake-style scalability and multi-format data storage. Some of the key technologies that make lakehouse operations possible are open table formats Delta Lake and Apache Iceberg and Apache Hudi to enable transactional storage in object storage, provide schema evolution and time travel features, and execute queries fast like warehouses. Performance results show significant improvements because, instead of 24 hours, Life Sciences applications now process their data in 30 minutes, which gives a 48 times better performance. A medallion architecture is one of the common ways to set up data in lakehouse systems, and it features three main layers: bronze, silver, and gold. The bronze layer provides the ability to ingest data in its raw form, retaining the characteristics of the source data. Further, the data is cleaned and validated by the silver layer into an analyzable state while ensuring consistency in formatting and the passing of quality metrics. Finally, the gold layer is where the data undergoes its final transformation for business consumption-usually by aggregating it in a way that makes sense from the point of view of the decision-making aspect. Interestingly, Microsoft implements this concept of bronze-silver-gold in the processing and tracking of origins with quality metrics throughout different stages of their Life Sciences data solutions [12].
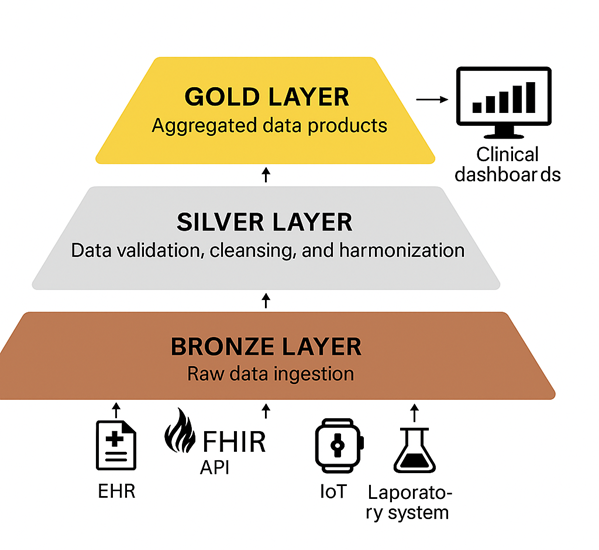
Figure 1: Medallion Architecture Layers in Life Sciences Data Lakehouse
This figure highlights how data quality and governance improve progressively across layers, ensuring traceability and readiness for AI-driven Life Sciences analytics.
Data Mesh and Domain-Oriented Architecture
Data mesh introduces an extensive architectural transformation apart from the centralized platform architectures found in warehouses and lakehouses. Data mesh functions according to four fundamental principles of domain-based decentralized data ownership and product-oriented data treatment and self-service infrastructure, and federated governance of computation. In the data mesh approach, all data management responsibilities are distributed among the teams of business areas, including radiology, pharmacy, and research engagement, rather than depending on a single platform team for data management. The approach treats datasets as essential products which require defined ownership and quality benchmarks and service level agreements while designers focus on user requirements. Automated tools are given to the domain teams through a self-service data infrastructure that allows the creation, publishing, and consumption of data products without requiring profound data engineering knowledge. Federated computational governance balances independence with necessary standardization through computational policies that automatically ensure security, privacy, quality, and interoperability standards.
Data mesh principles acknowledge that, in Life Sciences contexts, different data needs, vocabularies, and expertise are required across various domains such as clinical, operational, financial, and research. The cardiology department of a biopharma is deeply knowledgeable in cardiovascular data, but has very little understanding of the structures used in oncology. Data mesh operates through domain ownership of data assets which maintains interoperability through standardized interface and metadata contract definitions. The Life Sciences data mesh market will expand to USD 8.98 billion by 2033 because organizations now understand that their current centralized systems lack the ability to meet the various requirements of complex Life Sciences organizations 14. Organizations need to implement major transformations which include cultural changes for product thinking and platform development expense management.
The architectural bond between lakehouse and data mesh functions as a unified system which operates in harmony instead of creating opposition between them. Lakehouses provide the technological capability to house and manage the data, while the data mesh provides the structural model that allows for the decentralized distribution of ownership and responsibility. Most organizations are using lakehouse infrastructure as the foundation for data mesh, whereby the lakehouse serves as the self-service data platform upon which domain teams build and manage their data products. This scales both in technology through performance of the lakehouse, but also in organization through shared ownership.
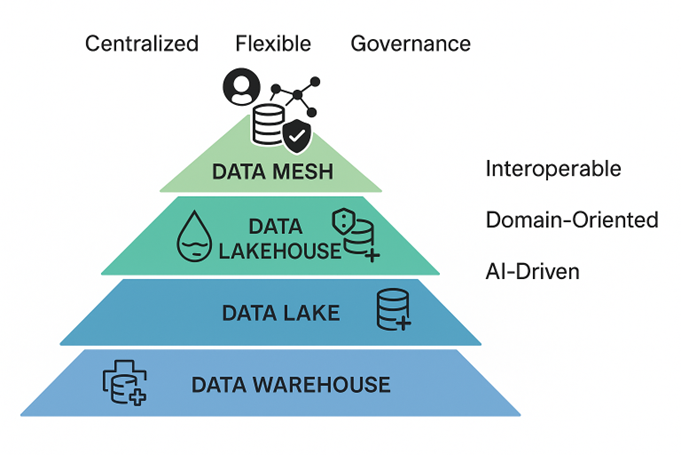
Figure 2: Evolution of Life Sciences Data Architectures – From Centralized Systems to Domain-Oriented Models
This figure highlights how technical scalability and organizational scalability converge to enable AI-driven and interoperable Life Sciences analytics.
Interoperability and FHIR Integration
Fast Life Sciences Interoperability Resources are the established standard for health data exchange and are integral for facilitating modern architectures. FHIR is a resource-oriented data model where specific resources reflect clinical concepts about clinical trials, observations, medications, and procedures with standardized schemas and relationships. The standard represents many state-of-the-art web technologies, including RESTful APIs, JSON and XML formats, and OAuth2 for authentication, making it much easier for developers to implement than earlier standards from Health Level Seven [15]. Implementation was accelerated by the mandate for FHIR APIs in the 21st Century Cures Act, and 84 percent of survey participants anticipate increased adoption of FHIR. In general, the integration of FHIR data in data lakehouse architecture is done in the bronze layer, which imports FHIR resources from source systems in their raw form for storage. The silver layer then transforms FHIR data into analytic formats, often in standard data models such as the Observational Medical Outcomes Partnership for longitudinal studies or maintains FHIR in its structure for operational analytics. This integrated FHIR data enables near real-time data streaming from the EHR systems for various use cases, including trial data harmonization, research collaboration, and cohort analytics.
The Life Sciences system has achieved major progress, but it still encounters major challenges with data sharing between different systems. The workforce lacks FHIR knowledge according to systematic reviews which show that more than 75% of surveyed participants identified knowledge deficits as their main challenge. The situation becomes worse because organizations use different methods to implement their systems and EHR vendors do not provide sufficient support and there are no standardized terms or value sets, and insufficient complex clinical test data exists for development purposes. The achievement of semantic interoperability depends on two essential factors which include FHIR syntax agreement and shared control of vocabularies and clinical procedures and data quality standards. The successful implementation of these elements needs continuous teamwork between standards organizations and software developers and Life Sciences providers.
Artificial Intelligence and Machine Learning Integration
The inclusion of artificial intelligence is the major driving element in modernizing data architecture. Modern data platforms should provide complete support for the entire machine learning lifecycle, from data preparation and feature engineering to model training, deployment, monitoring, and retraining. The foundation for implementing the concepts of automation, modularity, version control, repeatability, and monitoring in these workflows is provided by MLOps architecture. While ensuring that the models deployed are reliable, traceable, and managed, well-designed MLOps architecture should allow for quick experimentation with features and model configurations.
Adoption has significantly increased, with 22% of firms reporting use of specialist AI technologies in 2024-nearly ten times as many as in 2023. Pharmaceutical and biotechnology organizations lead adoption at 27%, followed by clinical research networks at 18%, and regulatory data platforms at 14%.. Spending on Life Sciences AI surged to USD 1.4 billion in 2024, roughly tripling from the prior year and well outpacing the combined total for the vertical AI market in all other industries. At this scale, investment proves that AI can solve several of Life Sciences' most intractable challenges: administrative burden reduction, hardening clinical decision support, and continuing to advance population health management through predictive analytics. Examples of specific AI applications now driving clinical value include automated clinical data abstraction, which reduces researcher documentation time by more than two hours daily; AI-powered trial-protocol approval systems that cut review times and data validation errors; predictive models for adverse event detection and protocol deviation forecasting that allow for earlier intervention; and natural language processing for automated annotation and metadata classification that increases accuracy and shortens the revenue cycle length. The Patient-Centered Analytical Learning Machine (PALM) developed at Montefiore demonstrates how a semantic data lake can integrate AI and ML at scale within regulated Life Sciences environments is a model of expansive AI deployment, centered around a semantic data lake framework that allows integration of AI and machine learning into operations without requiring wholesale replacement of infrastructure.
RQ1: Architectural Differences and Life Sciences Implementations
The idea that such a change in architecture—from conventional data lakes and warehouses to lakehouses and contemporary data mesh frameworks—represents a necessary transformation in Life Sciences data management is supported by this investigation. Although the traditional data warehouse provided secure, schematized storage with strong integrity and analytic performance, its use was limited in handling unstructured or semi-structured clinical data, such as image files, genomics data, and device streams. It is out of these limitations that the data lakes were born-supportive of varied types of data and schema-on-read flexibility. However, when strict governance and active metadata management are lacking, numerous Life Sciences data lakes have grown into disorganized, siloed repositories commonly known as data swamps, in which quality and discoverability are lost.
Lakehouse architecture bridged this gap by combining the scaling and flexibility of data lakes with governance, schema enforcement, and ACID transaction support of data warehouses. Implementations based on open-table formats like Delta Lake and Apache Iceberg have shown that it is now possible to achieve warehouse-grade performance on low-cost object storage by Life Sciences organizations. The adoption of medallion-layer structures, such as bronze, silver, and gold, further ensures traceability and quality progression across data pipelines. While the data mesh represents more an organizational than a purely technical evolution, ownership in data becomes distributed by alignment to the specific Life Sciences domains: radiology, oncology, and research engagement. Innovation is accelerated, while accountability is brought about. Complementary rather than competing, the architecture of mesh and lakehouse offer distributed ownership and governance through mesh, while lakehouse provides the technical foundation.
RQ2: Adoption Trends and Key Drivers
In fact, new data architectures have been adopted at an accelerating pace among Life Sciences organizations worldwide between 2019 and 2025. Today, an estimated seventy percent of new analytics projects in Life Sciences use lakehouse capabilities. This tells a clear story-the direction of travel in the Life Sciences sector toward unified, adaptable, and AI-enabled data environments. Four major factors are driving growth today: AI and predictive analytics, interoperability needs, especially regulatory frameworks like FHIR and the 21st Century Cures Act, the need for real-time analytics to drive operational efficiency, and lastly, cloud-based infrastructures are being adopted to partially offset the implementation challenges. The market of Life Sciences Data Mesh will grow from approximately $1.82 billion in 2024 to nearly $9 billion by the end of 2033, growing at an annual growth rate of more than nineteen percent. This transformation is not merely technological in nature; it's strategic, lying at the core of fulfilling the ever-growing needs of the Life Sciences industry for insight-informed decisions, personalized medicine, and good governance.
RQ3: Technical, Organizational, and Regulatory Challenges
This, however, reflects that some of the implementation hurdles remain. More specifically, one of the main ways in which this drive for improvement is still limited pertains to interoperability of technologies. Having a lot of formats and vocabularies out there, fragmented further by variations among providers of electronic health records, creates barriers for open information exchange even with the advent of the FHIR standard. The challenges in integrating legacy systems would apply quite naturally. Most of the biopharma research networks still use old on-premises systems incapable of supporting near-real-time data streaming.
The organizational obstacles include major cultural shifts: these companies need to move from a centralized data engineering team to domain-driven product ownership, which means extensive training and new governance models. This is relatively hard to implement in some regions that have very strict rules, for example, HIPAA and GDPR. Architectural renovation is really viable only if it is matched by organizational readiness, clear legislation, and people development.
RQ4: FHIR Integration and Semantic Interoperability
Fast Life Sciences Interoperability Resources (FHIR) serves to bridge modern data architectures with clinical systems. The literature review has confirmed that FHIR is now the dominant framework for Life Sciences data exchange, wherein over eighty percent of organizations plan to extend its use. FHIR data in modern lakehouse implementations often undergo ingestion at the bronze layer in native format and move into transformation at either the silver or gold layers into standardized models for analytics. This layered approach allows for regulatory traceability with high-performance querying at the same time for clinical analytics. However, semantic interoperability, which means arriving at a common meaning across systems, remains unresolved. Though FHIR brought syntactic standardization, the differences within terminologies, implementation profiles, and poor-quality data further reduce the effectiveness of FHIR. As partial workforce knowledge and only partial compliance of vendors further limit its full potential, achieving true semantic interoperability will require consistent implementation of standardized vocabularies, clinically complex test data development, and collaboration on governance across institutions.
RQ5: Integration of Artificial Intelligence and Machine Learning
Artificial intelligence has emerged as both a driver and a recipient of modern data architecture. Lakehouse and mesh empower MLOps pipelines that enable automation of feature engineering, model training, validation, deployment, and monitoring-all functionalities that conventional systems cannot provide. The research estimates that from 2023 to 2025, Life Sciences AI adoption increased tenfold, with approximately twenty-two percent of Life Sciences organizations deploying domain-specific AI tools and cumulative spending reaching USD 1.4 billion in 2024. Both AI and modern data architecture feed each other: a contemporary infrastructure cuts down hours in model building-to-deployment cycles to mere minutes and ascertains the availability of predictive insights right at the point of care. Real-world examples include sepsis prediction, clinical documentation automation, AI-driven prior authorization, and diagnostic imaging analysis. All these outcomes confirm that AI's success in Life Sciences will never be divorced from the robustness, scalability, and interoperability of the underlying data architecture.
RQ6: Performance Improvements and Clinical Outcomes
Several recent studies demonstrate the advantages of the lakehouse system in Life Sciences. For example, it has reduced data refresh intervals to below thirty minutes compared to a whole day and can perform queries almost twice as fast. A better data monitoring system with minimal errors helps reduce compliance issues. This means research teams can respond to findings much quicker, improving trial timelines and data reliability, and there is much better coordination in their care. These changes in handling data mean there is improved reliability of research data and significant improvement in the quality of analytical insights generated.
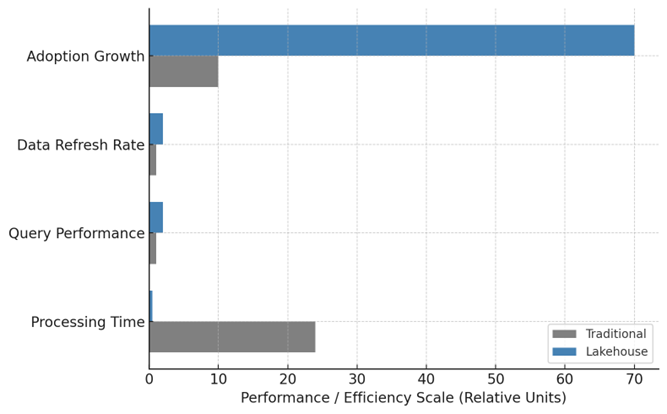
Figure 3: Quantitative Impact of Modern Data Architectures on Life Sciences Performance
This bar chart compares Traditional and Lakehouse architecture across four key Life Sciences performance metrics — Processing Time, Query Performance, Data Refresh Rate, and Adoption Growth.
RQ7: Best Practices and Strategic Recommendations
In updating data, several best practices should be considered by an organization. First, there is a need for a phased strategy wherein an organization starts with the most value-laden initiatives and scales efforts across the entire organization. Of great importance, too, is effective governance, striking a balance between centralized policies and the autonomy of different teams. For workforce shortages, organizations can institute training in FHIR standards, the creation of data products, and cloud design. Architecture must be ready for AI, including reproducibility and oversight. The voice of the clinical, operational, and technical teams should be included in making architecture decisions to align and truly improve clinical trial care while supporting workflows.
Key Findings Synthesis
The main points that came to the fore from this analysis that we feel are important to highlight include the following: First, the architecture of the data lakehouse closes the gap in agility and speed between the traditional data warehouses and data lakes while offering robust support for transactions, maintaining integrity in data structure, and ensuring effective governance that is scalable and can handle a variety of data formats. Most importantly, performance improvements can be up to double the speed, refreshing data in minutes, not hours. That has considerably enhanced efficiency and is major in facilitating timely decision-making in research and development settings. A bronze, silver, and gold medallion framework clearly shows ways in which data will be organized in its transformation from raw collection to validated upgrades, with improvements assured to be implemented and tracked at its source.
Third, principles guiding domain-oriented ownership for data-as-products within a data mesh address issues about organizational scalability directly and in ways that no purely technical solution could. Shared ownership allows leverage of domain expertise while achieving interoperability through federated government. Implementation requires significant management of organizational change and cultural change and deployment of a technical platform. Fourth, FHIR will be key for leveraging data from electronic health record systems, but semantic interoperability requires sustained attention to workforce knowledge development, consistency of vendor implementation, and standardization of terminologies with collaborative governance. Fifth, AI is both a major driver and major beneficiary of modernizing data architecture: well-designed infrastructure allows the automation, reproducibility, and governance that are needed for secure, productive large-scale deployment of regulated Life Sciences AI.
Advantages and Disadvantages
The benefits of modern architecture are that data lakehouse and data mesh models have some key advantages when compared to traditional architecture. High performance: Both models support analytics in research and operational workflows either in real time or close to it. They unify data on one platform and prevent data silos, which helps in avoiding many redundant systems; schema flexibility allows for changes in sources without extensive remodeling. ACID compliance delivers consistency in regulated Life Sciences applications. Deployment, which is cloud-native, offers reduced expenses and flexible scaling. Vendor lock-in is avoided with open table formats. Regulatory compliance is made possible by enhanced governance and lineage monitoring. All operations, from business intelligence to machine learning, are made simpler and less complex by native support for a broad variety of analytics workloads.
The new Life Sciences data system raises many issues from several standpoints. It would be very expensive: modernizing the system may cost the country tens of billions of dollars. In practice, a data mesh approach might not be as fluid as transition to. Organizational change is complex to establish as many workers may not be familiar with modern technologies and standards like FHIR. In addition, incorporating those new tools into the organization is tough, as that has to align with old systems, keeping technical debt in mind. Beyond that, the potential to make a variety of organizations communicate smoothly in this type of framework entails an immense amount of coordination and continuous efforts even after standards are developed. In general, managing a distributed architecture adds more complexity. Such adaptation of those systems calls for specialized knowledge. There's always a risk, too, during the migration phase, which could end up losing data and/or disrupting services–which nobody wants. Lastly, most of the emerging technologies are still evolving, and best practices are limited compared to established systems.
Implications for Practice
Life Sciences and biopharma organizations need to renew their data infrastructure strategically, beginning with use cases yielding high impact, clear benefits, and returns on investment. This needs to go hand in hand with enhancing the competencies of the people in the organization. Any investment made in workforce development should therefore be targeted at training programs based on current data technologies such as FHIR standards, cloud systems, and data product strategies. Organizations need to set up a strong data governance framework prior to delving deep into more complex, distributed systems. This saves them from chaos later on. Interoperability standards mainly will be critical to integrating operational data effectively; this means mostly FHIR. Design for an AI future state as early as possible-even if the higher-order uses of AI are not included, it ensures the relevance of the future investment. What's important in every stage is participation by all those involved in the research and operational processes to make sure changes align with actual workflow needs. Lastly, improvement of an organization's data architecture is a multi-year process. It is critical that the leadership continues to invest and provides sustained support rather than considering the effort a single project.
This recent analysis underlines the very significant shift now underway in Life Sciences data architecture-from the older model of centralized data lakes and warehouses to more modern forms, lakehouses, and decentralized approaches to data mesh. It could be game-changing for how we use Life Sciences data in research, clinical development, and translational analytics applications, and it is not going at a snail's pace. Indeed, by 2025, it is estimated that 70 percent of new Life Sciences data projects will contain lakehouse features. Meanwhile, with increased awareness of the shortfalls of the traditional model and the appeal of newer solutions, the market for Life Sciences data mesh is set to reach almost $9 billion by 2033. It also underlines a number of key gaps in current research: long-term studies are needed for return on investment, total cost of ownership, and the impact of different architectural models on research efficiency and trial outcomes. What it means is that while we have good data about performance and adoption trends, we can tell very little regarding cost-effectiveness and research benefit. This, in our view, is where we would gain more clearer insights through controlled studies comparing organizations pursuing different strategies. Evidence such as this will support more-informed decision-making in the years ahead.
There is a need for further research on effective practices in managing organizational change associated with Life Sciences data architecture transformation. Although the technical literature is quite well advanced, the organizational, cultural, and workforce development aspects are less so. Case studies of successful and unsuccessful implementations, if supported with detailed analysis of organizational factors leading to each outcome, would be beneficial. Finally, we need to study the federated governance frameworks that would balance domain autonomy with central oversight. Unique governance needs in Life Sciences driven by regulatory challenges and requirements for patient safety are not fully addressed in general enterprise data mesh literature. Fourth, in the context of increasing concerns about the protection of clinical trial data, research related to privacy-preserving technologies and their interaction with modern data architecture is of growing importance. Techniques that appear promising include federated learning, homomorphic encryption, differential privacy, and secure multi-party computation; all these will require further study regarding implementation issues in practical Life Sciences settings. Fifth, exploration of edge computing integration with centralized data platforms for real-time processing of data from Internet of Things medical devices and wearables represents an emerging frontier. Systematic investigation of the architectural implications of hybrid edge-cloud processing for Life Sciences applications will be warranted.
Sixth, the development of standardized data product templates, reference architectures, and implementation patterns for common Life Sciences domains such as imaging, laboratory, pharmacy, and care management would drive adoption by reducing the implementation effort. The creation of an open-source repository of Life Sciences data product patterns would offer value to a wide community. Finally, investigation of the implications of artificial intelligence foundation models and large language models on Life Sciences data architecture is required. The unique demands introduced by these models in terms of training data curation, fine-tuning infrastructure, inference serving, and responsible AI monitoring require architectural capabilities unlike those from traditional analytics alone and thus deserve focused research attention. Collectively, these research directions would advance the theoretical understanding and practical implementation of modern Life Sciences data architecture.
REFERENCE

Sumanth Singh, From Data Lakes to Data Products: The Future of Enterprise Data Architecture in Life Sciences, Int. J. of Pharm. Sci., 2025, Vol 3, Issue 11, 2886-2900. https://doi.org/10.5281/zenodo.17649479
 10.5281/zenodo.17649479
10.5281/zenodo.17649479