
We use cookies to ensure our website works properly and to personalise your experience. Cookies policy

We use cookies to ensure our website works properly and to personalise your experience. Cookies policy
Adhiparasakthi College of Pharmacy, Melmaruvathur, Tamilnadu-603 319.
In the present study, a series of Pyrazolidine 3, 5 Dione derivative compounds were evaluated for their molecular binding affinity using in silico docking techniques. Docking simulations were performed with ArgusLab and PyRx software to investigate the interaction of the ligands with the target protein. The binding energies obtained from both ArgusLab and PyRx were compared with those of standard drug (Indomethacin). Results revealed that the Pyrazolidine 3, 5 Dione derivative compounds shows better (more favorable) binding energies than Indomethacin in both ArgusLab and PyRx analyses, indicating stronger hydrogen bonding and more stable binding interaction. These findings suggest that the Pyrazolidine 3, 5 Dione hold promise as potential lead molecules for Anti-inflammatory drug development.
Inflammation is the immune system’s response to harmful stimuli, such as pathogens, damaged cells, toxic compounds, or irradiation, acts by removing injuries stimuli and initiating the healing process. Inflammation is therefore a defence mechanism that is vital to health. At the tissue level, inflammation is characterized by redness, swelling, heat, pain and loss of tissue function, which result from local immune, vascular and inflammatory cell responses to infection or injury. Various pathogenic factors, such as infection, tissue injury, or cardiac infarction, can induce inflammation by causing tissue damage. Cyclooxygenase also abbreviated as COX, is a prostaglandin endoperoxide synthase, enzyme involved in the metabolism of Arachidonic acid (AA) and synthesis of prostanoid proinflammatory prostaglandins (PGE2, PGF2α). Nonsteroidal anti-inflammatory drugs (NSAIDs) represent one of the most common classes of medications used worldwide with an estimated usage of >30 million per day for inflammation and related disorder. Most of the NSAIDs are carboxylic acid containing drugs including salicylate derivatives (aspirin), carboxylic and heterocyclic acid derivatives (indomethacin), propionic acid derivatives (ibuprofen, ketoprofen). These organic acid containing drugs act at the active site of the enzyme preventing the access of Arachidonic acid (AA) to the enzyme and stop the cyclooxygenase pathway. Unfortunately, besides the excellent anti-inflammatory potential of the NSAIDs, the severe side effects such as gastrointestinal (GI), ulceration, perforation, obstruction and bleeding has limited the therapeutic usage of NSAIDs. Inflammation is an intrinsically beneficial event that leads to removal of offending factors and restoration of tissue structure and physiological function. This is consistent with the idea that inhibition of COX-1 underlies the gastrointestinal side effects.
The Immune Response
The immune response occurs when immunologically competent cells are activated in response to foreign organisms or antigenic substances liberated during the acute or chronic inflammatory response. The outcome of the immune response for the host may be beneficial, as when it causes invading organisms to be phagocytosed or neutralized. On the other hand, the outcome may be deleterious if it leads to chronic inflammation without resolution of the underlying injurious process. Chronic inflammation involves the release of number of mediators that are not prominent in the acute response. Chronic inflammation results in pain and destruction of bone and cartilage that can lead to severe disability and in which systemic changes occur that can result in shortening of life. The cell damage associated with inflammation acts on cell membranes to cause leukocytes to release lysosomal enzymes; Arachidonic acid is then liberated from precursor compounds.
The Components of the Acute Inflammatory Reaction:
The acute inflammatory reaction has two components. An innate, non-adaptive response, thought to have been developed early in evolution and present in some form or other in most multicellular organisms.
The Innate Immune Response:
The innate response occurs immediately on injury or infection. It comprises vascular and cellular elements. Mediators generated by cells or from plasma modify and regulate the magnitude of the response. Tissue macrophages, bearing toll receptors, recognize specific pathogen associated molecular patterns on the microorganism and release cytokines, particularly interleukin (IL)-1 and tumour necrosis factor (TNF)-α as well as various chemokines.
Exudate contains enzyme cascades that generate bradykinin, and C5a and C3a. Complement activation lyses bacteria. C5a and C3a stimulate mast cell to release histamine which dilates local arterioles. Tissue damage and cytokines release prostaglandin and leukotriene. Cytokine stimulate synthesis of vasodilator nitric oxide, which increase vascular permeability. Using adhesion molecules, leucocytes roll on, adhere to and finally migrate through vascular endothelium towards the pathogen, where phagocytosis and killing takes place.
The Adaptive Response:
The adaptive immunological response boosts the effectiveness of the innate responses. It has two phases,
Antibodies provide:
Cell- mediated reactions involve:
Systemic Responses in Inflammation:
In addition to the local changes in an inflammatory area, there are often general systemic manifestations of inflammatory disease, including fever, an increase in blood leucocytes termed leucocytosis, and the release from the liver of acute phase proteins. These include C-reactive protein, α2- macroglobulin, fibrinogen, α1-antitrypsin and some complement components. While the function of the many components is still a matter of conjucture, they all seem to have antimicrobial action. C-reactive protein, for example, binds to some microorganisms, and the resulting complex activates complement.
Unwanted Inflammatory and Immune Responses:
The immune response has to strike a delicate balance. According to one school of thought, an infection-proof immune system would be a possibility but would come at a serious cost to the host. With approximately 1 trillion potential antigenic sites in the host, such a super immune system would be some 1000 times more likely to attack the host itself, triggering autoimmune disease. In practice, therefore it is not uncommon to find that innocuous substances such as pollen, or the hosts own tissues, sometimes inadvertently activate the immune system; when this occurs, anti-inflammatory or immunosuppressive therapy may be required. Unwanted immune responses, termed allergic or hypersensitivity reactions, have been classified in to four types.
Mechanism:[1]
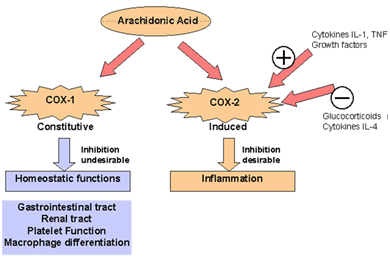
Figure 1: Mechanism Of Action
Inflammation is a complex biochemical mechanism which leads to the stimulation of infection agents and promotes injury. It causes tissue damage and pain monitored through treatment. Chemical agents are secreted by immune cells like cytokines, chemokine and reactive oxygen species at injury site to eliminate pathogens. A major constituent of inflammatory process is arachidonic acid which is a byproduct of fast acting cell membrane. Arachidonic acid is transformed into prostaglandins and thromboxane enzyme by cyclooxygenase (COX). Neutrophils begin to attach strongly to the endothelium by using carbohydrate ligands to show symptoms of inflammation. Endothelial cell in their stimulated form is responsible for the production of surface bonded and soluble particles. They produce a strong adhesion between neutrophils and endothelium; Neutrophils leave the bloodstream and travels across endothelium. Production of particular cells like cell adhesion molecules (CAMs), their activators and chemical stimulus is responsible for the neutrophil’s emigration. Stimulation of these chemical agents initiates the formation of inflammatory cytokines including TNF, IL-1, chemokines and IL-6 that causes tissue damage. Due to phagocytic activity of cells, migratory neutrophils are eventually removed from inflammatory site through apoptosis and produce Anti-inflammatory cytokines. The inflammatory is a common pathogenesis of many chronic diseases, including cardiovascular and bowel diseases, diabetes, arthritis and cancer. Cyclooxygenase (COX) inhibition and the resulting inhibition of prostaglandin and other eicosanoid synthesis mitigate pain, fever, and inflammation. NSAID’s relieve fever and pain by inhibiting COX- mediated prostaglandin synthesis. The cyclooxygenase (COX) enzyme also known as prostaglandin endoperoxide H synthase (PGHS) exits in two isozymes PGHS-1 or COX-1 and PGHS-2 or COX-2. NSAID’s has helped in understanding the tissue mechanism of inflammation. It is generally agreed that the NSAID’s and analgesic drugs available in the market such as (aspirin, phenylbutazone, oxyphenbutazone, indomethacin and ibuprofen).
General Structure of Pyrazolidine 3, 5 Dione:[7]

Figure 2: Structure Of Pyrazolidine 3, 5-Dione
Therapeutic Action:
Pyrazolidine 3, 5-dione derivatives have a wide range of biological and pharmacological activities and play an important role in the sub-structures of various drugs. Pyrazolidine 3,5 diones exhibits miticidal, acaricidal, insecticidal, herbicidal, antimicrobial, anti-pyretic, anti-inflammatory activity, angiotensin receptor antagonists, antioxidant, anti-depressant, anti-diabetic, anti-Alzheimer and anticancer. The chemistry of pyrazolidine 3, 5-dione and their fused heterocyclic derivatives have received considerable attention owing to their synthetic and effective biological importance. In spite of a large number of antibiotics and chemotherapeutics available for medicinal use, the treatment of infectious diseases still remains an important and challenging problem because of a combination of factors including emerging infectious diseases and the increasing number of multi-drug-resistant microbial pathogens. Through the various molecules designed and synthesized for this aim, in recent years, active research has been initiated on heterocyclic and the chemistry of pyrazolidine-3,5-dione has received considerable attention owing to their synthetic and effective biological importance. Pyrazolidine-3, 5-dione moiety has been incorporated into a wide variety of therapeutically interesting into a wide variety of therapeutically interesting drug candidates including anti-inflammatory and anti-microbial. There are many marketing drugs containing the pyrazolidine 3, 5-dione group e.g., feprazone, phenylbutazone, indomethacin from the literatures, it may be predicted that pyrazolidine 3, 5-dione moiety represents important pharmacophore and plays a vital role in medicinal agents. In the design of new drugs, the combination of different pharmacophores may lead to compounds with interesting biological profiles.
Methods And Materials: [2,5,9,10]
Software’s Used
Arguslab
ArgusLab 4.0 is a user-friendly molecular modeling package which can be used for introductory modeling lesson in academics. Flexible ligand docking is possible with ArgusLab by describing by ligand torsions tree. The binding site of the protein is described by grids Ligand’s root node i.e. groups of bonded atoms that do not have rotable bonds, is placed on the grid in the binding site and different energetically favorable conformers are created by rotation. Each pose is scored using the scoring functions employed. N lowest energy poses are retained and this undergoes minimization, Re-clustering and ranking.
PyRx
Docking of all the ligands was performed with PyRx software and the highest binding energy (most negative) was recorded as the binding affinity. Visual analysis of the docking site was performed with discovery studio software.
Ligand Preparation:
Ligands can be retrieved from several databases such as ZINC, PubChem or can be sketched by applying the Chemsketch tool. Ligand 2D structures were drawn using ACD Chemsketch (freeware) 2015 2.5. Chem 3D viewer was used to convert the 2D structure into 3D. All the structures were saved as the pdb file format for input to PyRx 0.8 and all the ligand structures were then saved in mol file format, for input into Argus lab.
Preparation Of Protein:
Three-dimensional structure of the protein should be retrieved from the RCSB protein data bank (PDB); afterward the retrieved structure should be pre-processed for removal of heteroatom then energy minimization was performed by using both Argus and Pyrex software and visualization was done by using Discovery studio 3.5.
Molecular Docking:
Ligand was docked against the protein and the interaction was analyzed by using PyRx 0.8 and Argus lab. For the docking of ligands into protein active site and to estimate the binding affinities of docked compounds software is used. The protein-ligand interaction plays a significant role in structural based designing. All the compounds have shown ≥5 hydrogen bond donors, ≥ 10 hydrogen bond acceptors, molecular mass≥500 Da, highly lipophilic in nature and molar refractivity between 40-130.The rule describes molecular properties imperative for drugs pharmacokinetics in the human body, including absorption, distribution, metabolism and excretion. The scoring function gives the score on the basis of best-docked ligand complexed is picked out.
Table: 1 Derived compounds and its IUPAC Name
|
S.No |
Derived Compounds |
Iupac Name |
|
1. |

|
1-[2-(1H-1,3-benzimidazol-2-yl) phenyl]-2-(4-nitrophenyl) pyrazolidine-3,5-dione
|
|
2. |

|
1-[2-(1H-1,3-benzimidazol-2-yl) phenyl]-2-(2-chlorophenyl) pyrazolidine-3,5-dione
|
|
3. |
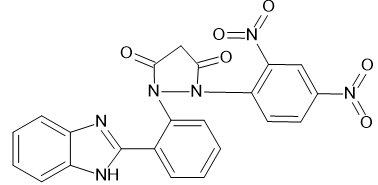
|
1-[2-(1H-1,3-benzimidazol-2-yl) phenyl]-2-(2,4-dinitrophenyl) pyrazolidine-3,5-dione
|
|
4. |

|
1-[2-(1H-1,3-benzimidazol-2-yl) phenyl]-2-(4-bromophenyl) pyrazolidine-3,5-dione
|
|
5. |

|
1-[2-(1H-1,3-benzimidazol-2-yl) phenyl]-2-(4-hydroxyphenyl) pyrazolidine-3,5-dione
|
|
6. |

|
1-[2-(1H-1,3-benzimidazol-2-yl) phenyl]-2-(3,4-dimethylphenyl) pyrazolidine-3,5-dione
|
|
7. |
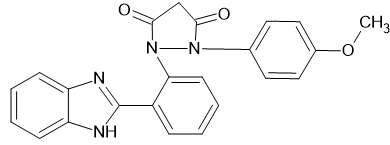
|
1-[2-(1H-1,3-benzimidazol-2-yl) phenyl]-2-(4-methoxyphenyl) pyrazolidine-3,5-dione
|
|
8. |
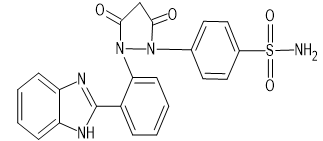
|
4-{2-[2-(1H-1,3-benzimidazol-2-yl) phenyl]-3,5-dioxopyrazolidin-1-yl} benzene-1-sulfonamide
|
|
9. |
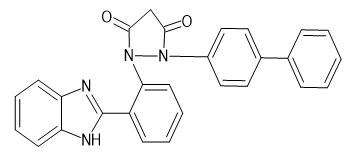
|
1-[2-(1H-1,3-benzimidazol-2-yl) phenyl]-2-(4-Biphenyl) pyrazolidine-3,5-dione
|
|
10. |
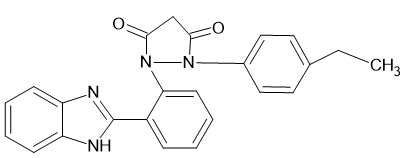
|
1-[2-(1H-1,3-benzimidazol-2-yl) phenyl]-2-(4-ethylphenyl) pyrazolidine-3,5-dione
|
RESULT AND DISCUSSION:
Swiss Adme:
Drug likeliness RADAR of compounds with good activity and ADME performed using the same procedure, and the binding affinity was obtained.
Table:2 Swiss Adme Results
|
Cmpd |
M.wt (g/mol) |
Number of heavy atoms |
Number of aromatic heavy atoms |
Fraction csp3 |
Number of rotable bonds |
Number of hydrogen bond acceptors |
Number of hydrogen bond donor |
Molar refractivity |
Tpsa (A?) |
Log po/w |
Log |
|
1 |
415.40 |
31 |
18 |
0.09 |
4 |
4 |
2 |
129.40 |
110.50 |
2.23 |
-3.63 |
|
2 |
404.85 |
29 |
18 |
0.09 |
3 |
2 |
2 |
125.59 |
64.68 |
2.85 |
-5.40 |
|
3 |
460.40 |
34 |
18 |
0.09 |
5 |
6 |
2 |
138.22 |
156.32 |
1.79 |
-4.94 |
|
4 |
449.30 |
29 |
18 |
0.09 |
3 |
2 |
2 |
128.28 |
64.68 |
3.09 |
-5.72 |
|
5 |
386.40 |
29 |
18 |
0.09 |
3 |
3 |
3 |
122.60 |
84.91 |
2.32 |
-4.67 |
|
6 |
398.46 |
30 |
18 |
0.17 |
3 |
2 |
2 |
130.51 |
64.68 |
2.95 |
-5.41 |
|
7 |
400.43 |
30 |
18 |
0.13 |
4 |
3 |
2 |
127.07 |
73.91 |
2.98 |
-4.88 |
|
8 |
449.48 |
32 |
18 |
0.09 |
4 |
5 |
3 |
131.57 |
133.22 |
2.10 |
-4.27 |
|
9 |
446.50 |
34 |
24 |
0.07 |
4 |
2 |
2 |
146.02 |
64.68 |
3.35 |
-6.29 |
|
10 |
398.46 |
30 |
18 |
0.17 |
4 |
2 |
2 |
130.35 |
64.68 |
2.97 |
-5.39 |
Tble:3
|
Compounds |
GI Absorption |
BBB Permeant |
PGP Substrate |
Cyp1a2 Inhibitor |
Cyp2c19 Inhibitor |
Cyp2c9 Inhibitor |
Cyp2d6 Inhibitor |
Cyp3a4 Inhibitor |
|
1 |
High |
No |
Yes |
No |
Yes |
Yes |
No |
No |
|
2 |
High |
Yes |
Yes |
No |
Yes |
Yes |
Yes |
Yes |
|
3 |
High |
No |
Yes |
No |
Yes |
Yes |
No |
No |
|
4 |
High |
Yes |
Yes |
No |
Yes |
Yes |
Yes |
Yes |
|
5 |
High |
No |
Yes |
No |
No |
Yes |
Yes |
No |
|
6 |
High |
Yes |
Yes |
No |
Yes |
Yes |
Yes |
Yes |
|
7 |
High |
No |
Yes |
No |
Yes |
Yes |
Yes |
Yes |
|
8 |
High |
No |
Yes |
No |
No |
No |
No |
No |
|
9 |
High |
Yes |
Yes |
No |
Yes |
Yes |
Yes |
No |
|
10 |
High |
Yes |
Yes |
No |
Yes |
Yes |
Yes |
Yes |
Docking Score:
The docking study was conducted using ArgusLab software against two target proteins 6COX and 1EQH. The pyrazolidine 3,5 dione derivative compounds exhibited better binding affinities (more negative docking score) compared to the standard drug. Some compounds showed significantly higher binding score such as -14.8772 and -13.667, indicating strong potential as COX inhibitors.
Table:4 Docking Score Result Using Arguslab Software
|
S.NO |
Structure |
Docking Score (Kcal/Mol) |
|
|
6COX |
1EQH |
||
|
STD |
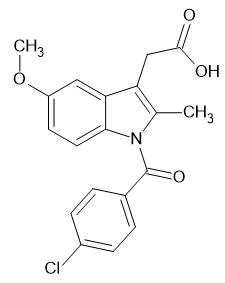
|
-12.372 |
-11.2349 |
|
1. |
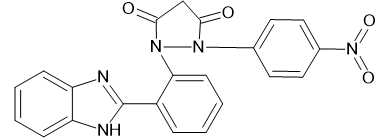
|
-9.35967 |
-10.6505 |
|
2. |

|
-10.0948 |
-12.3073 |
|
3. |
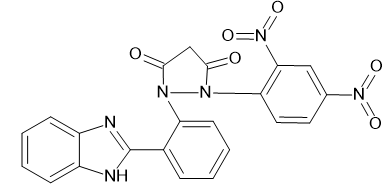
|
-10.4623 |
-10.6461 |
|
4. |
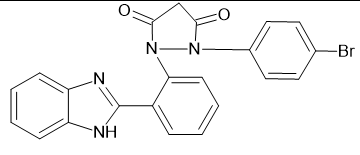
|
-9.39262 |
-12.3179 |
|
5. |
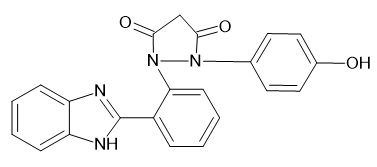
|
-9.12018 |
-11.6336 |
|
6. |

|
-14.8772 |
-12.5008 |
|
7. |

|
-11.6859 |
-11.4557 |
|
8. |

|
-9.6271 |
-10.6181 |
|
9. |
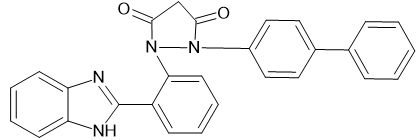
|
-11.5647 |
-13.4083 |
|
10. |
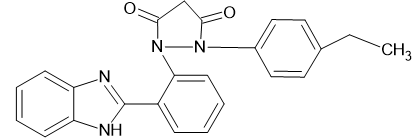
|
-13.6676 |
-13.3896 |
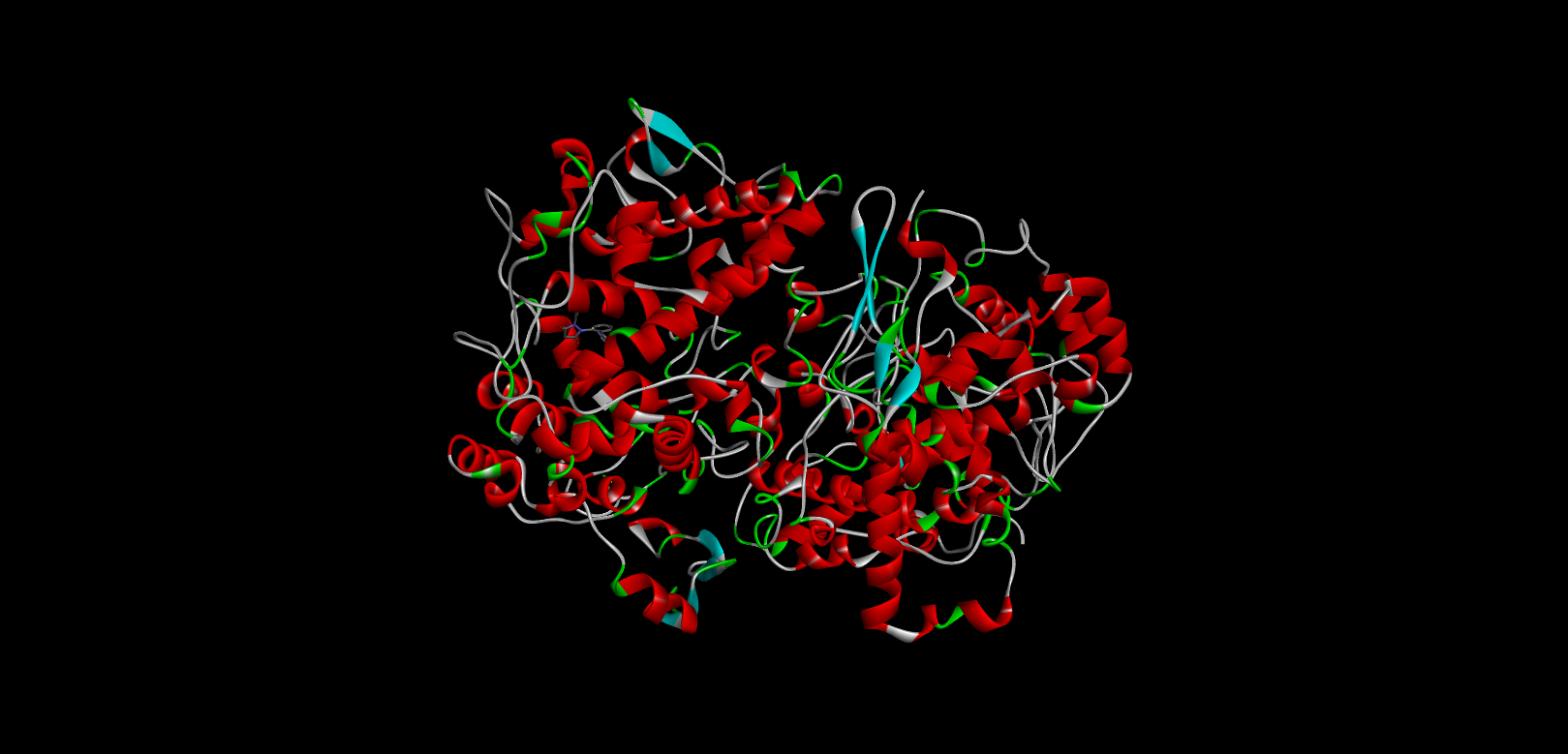
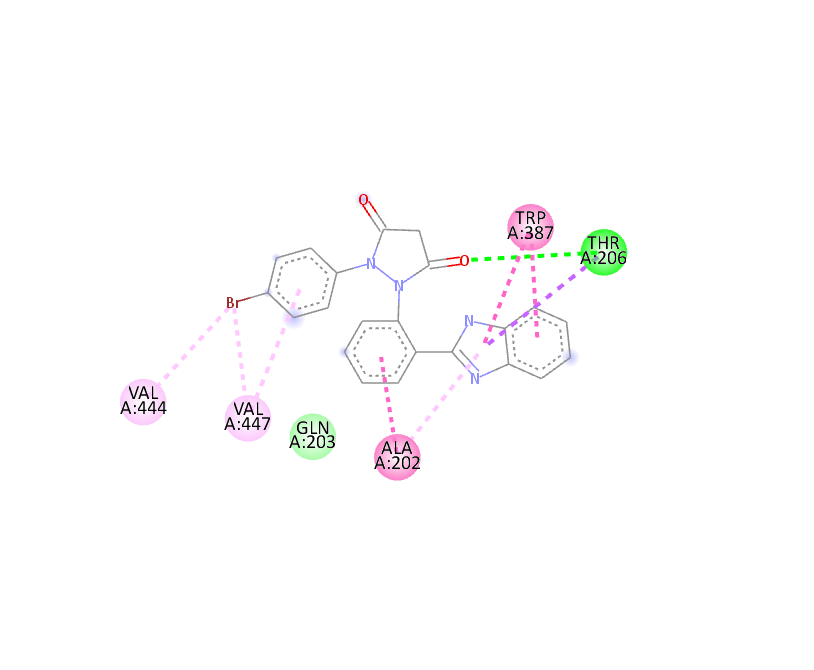
Docking Using Pyrx Software:
The docking study was conducted using PyRx software against two proteins, 6COX and 1EQH. The pyrazolidine 3,5 dione derivative compounds exhibited stronger binding affinity (more negative values) than the standard drug. The best binding score for 1EQH and 6COX was -10.4 and -11.9.
Table:5 Docking Score (6COX) Using PyRx Software
|
S. No |
Structure
|
Docking Image |
2d Image |
Docking Score |
|
Std 1 |

|
|
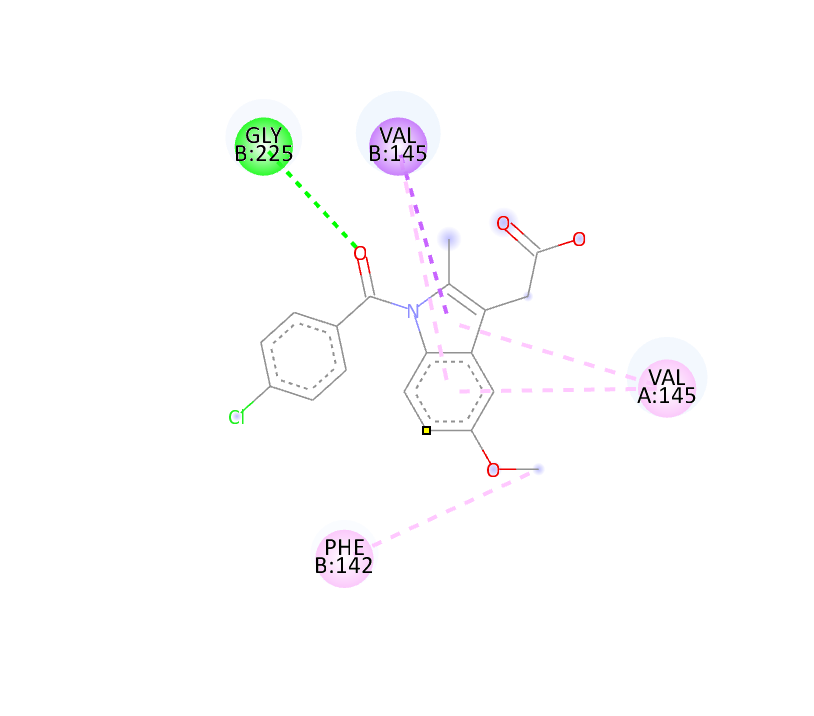
|
-8.1 |
|
|
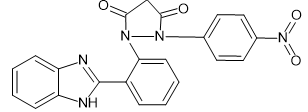
|
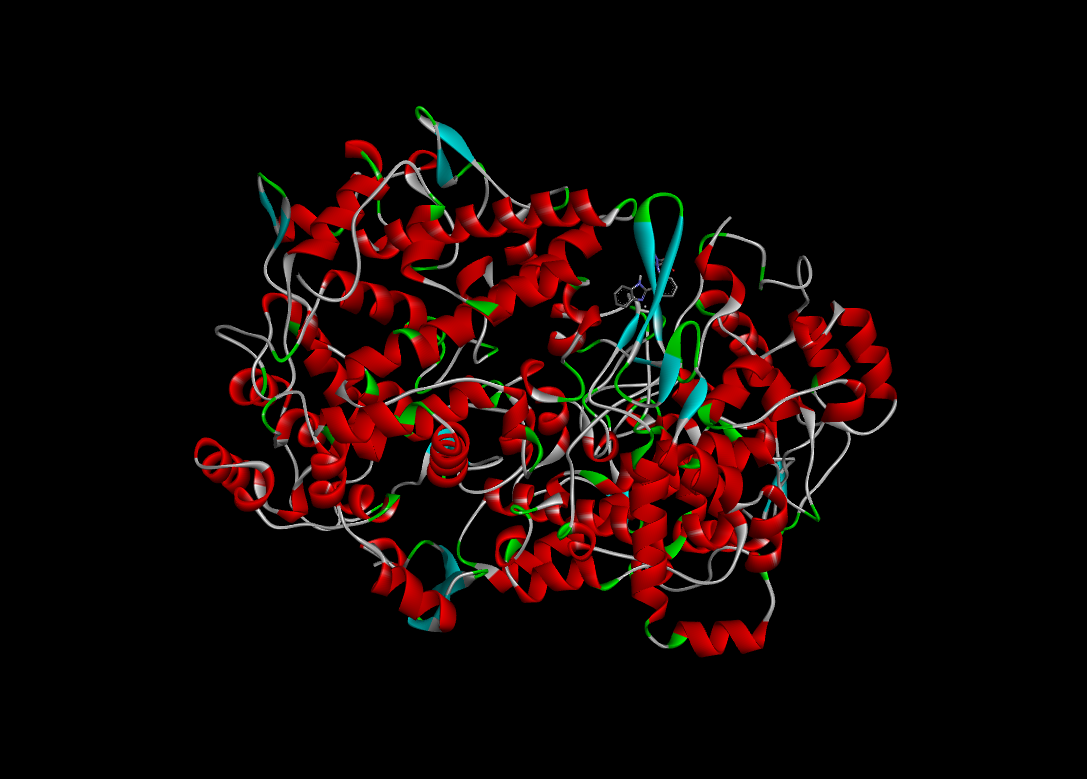
|
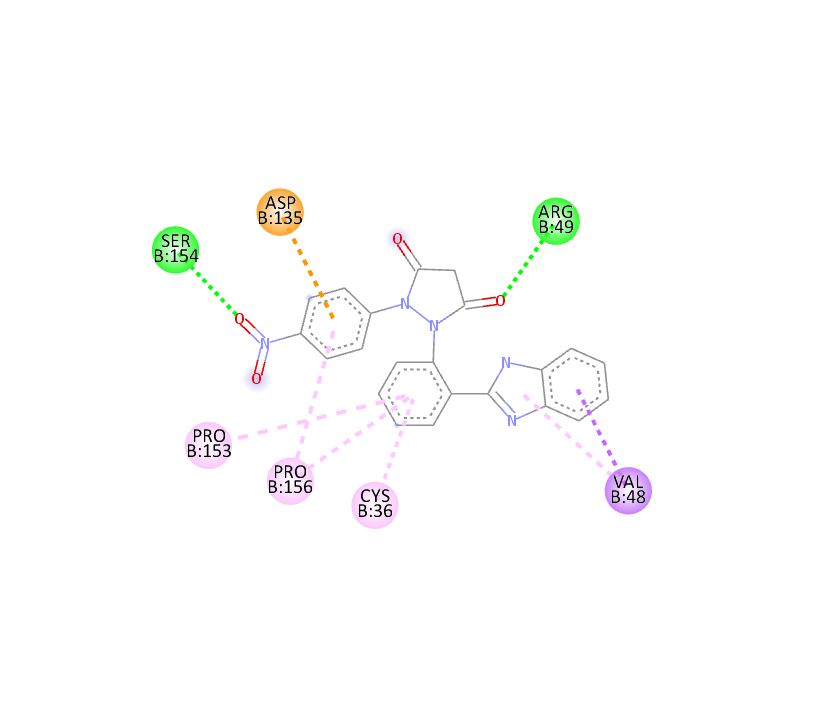
|
-10.2 |
|
|
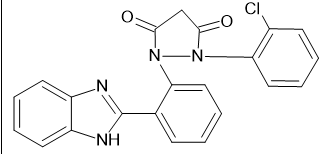
|
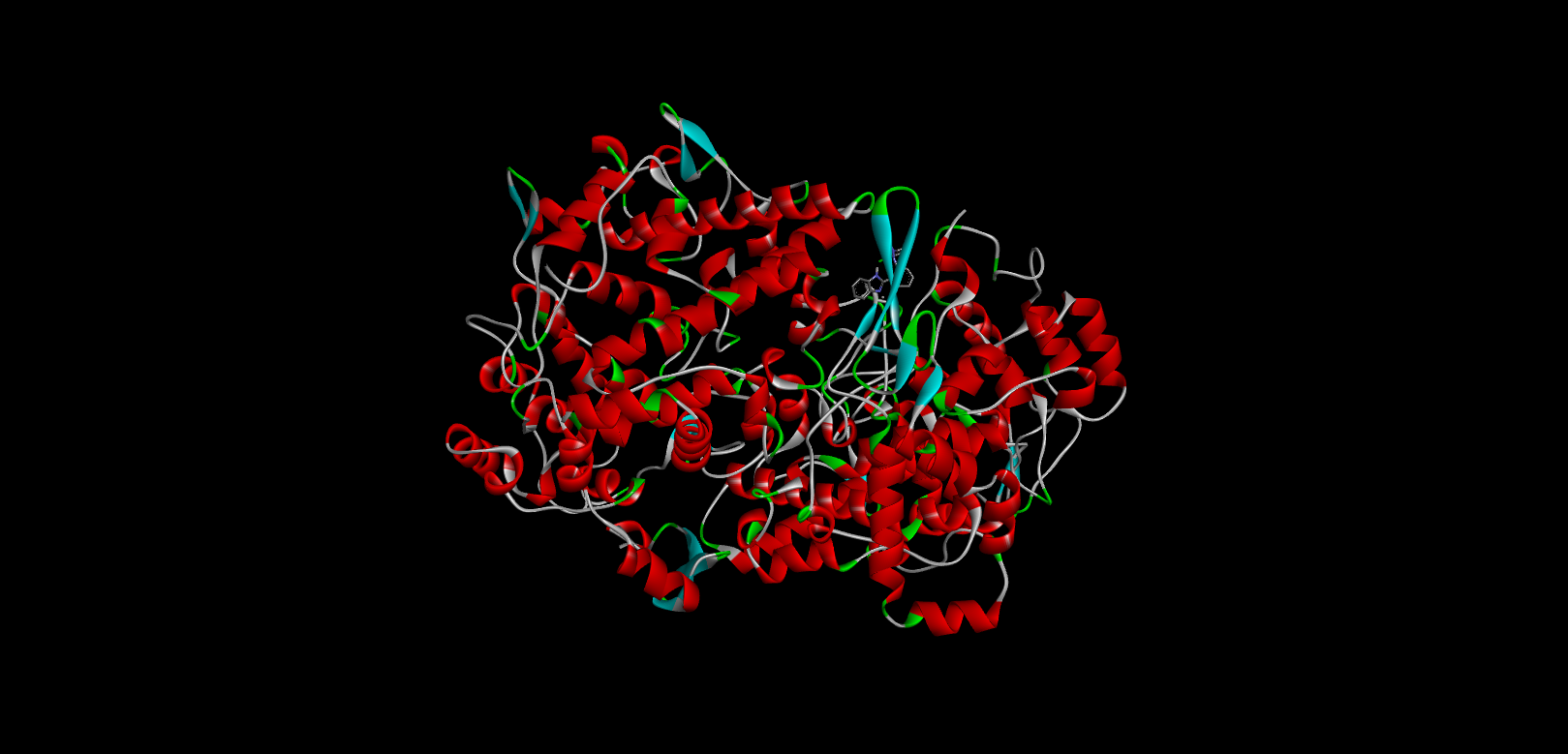
|
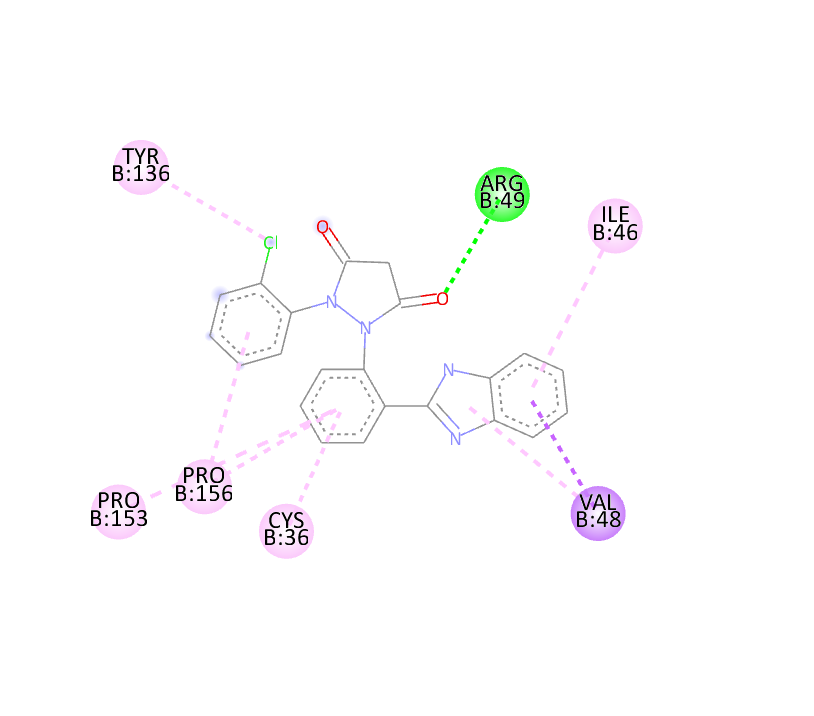
|
-10 |
|
|

|
|

|
-10.4 |
|
|
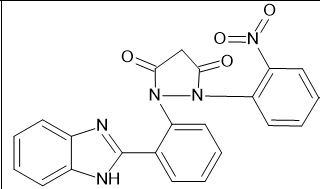
|
|
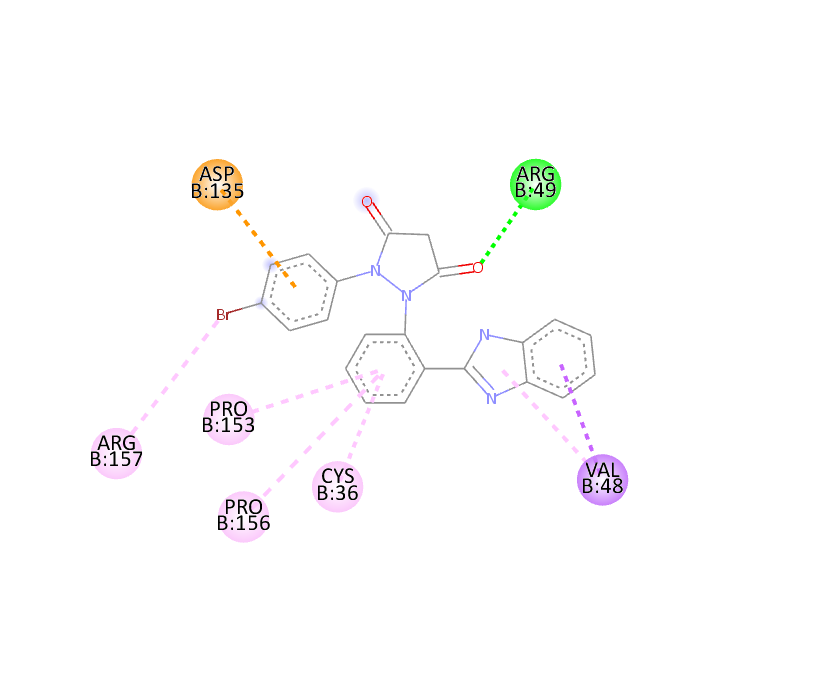
|
-9.9 |
|
|
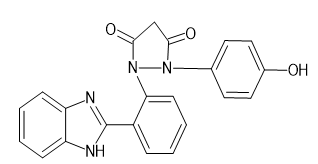
|

|
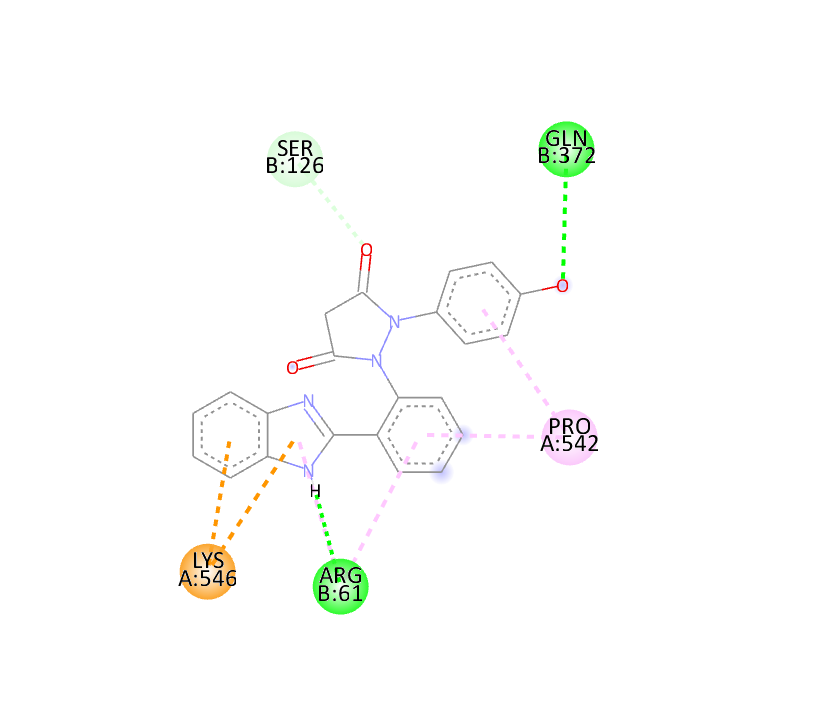
|
-9.9 |
|
|

|
|
|
-10.4 |
|
|

|
|

|
-10.1 |
|
|
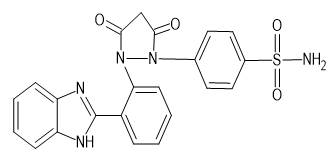
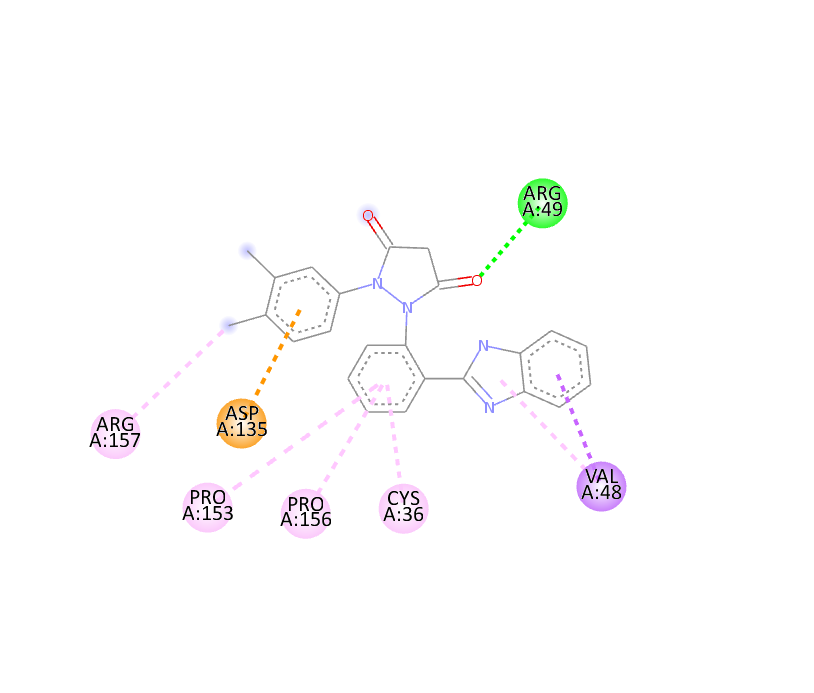
|
|
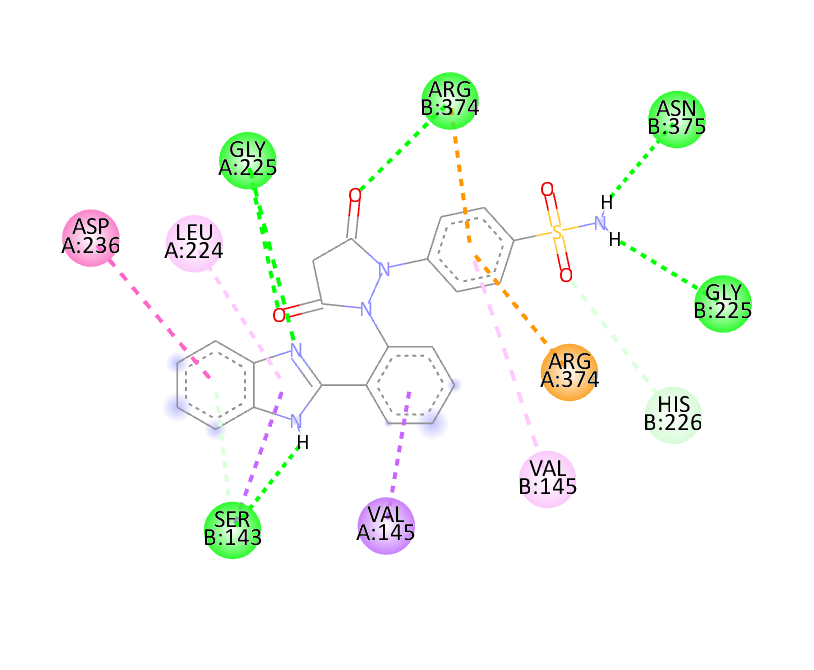
|
-10.2 |
|
|
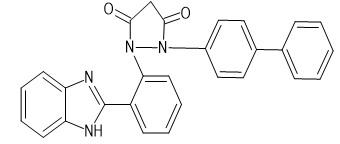
|

|

|
-9.8 |
|
|

|
|

|
-9.8 |
Table :6 Docking Score (1eqh) Using Pyrx Software
|
S. No |
Structure
|
Docking Image |
2d Image |
Docking Score |
|
Std 1 |
|
|

|
-8.5 |
|
|
|

|

|
-10.2 |
|
|
|
|

|
-10.9 |
|
|
|

|

|
-11.6 |
|
|
|

|
|
-11.9
|
|
|
|
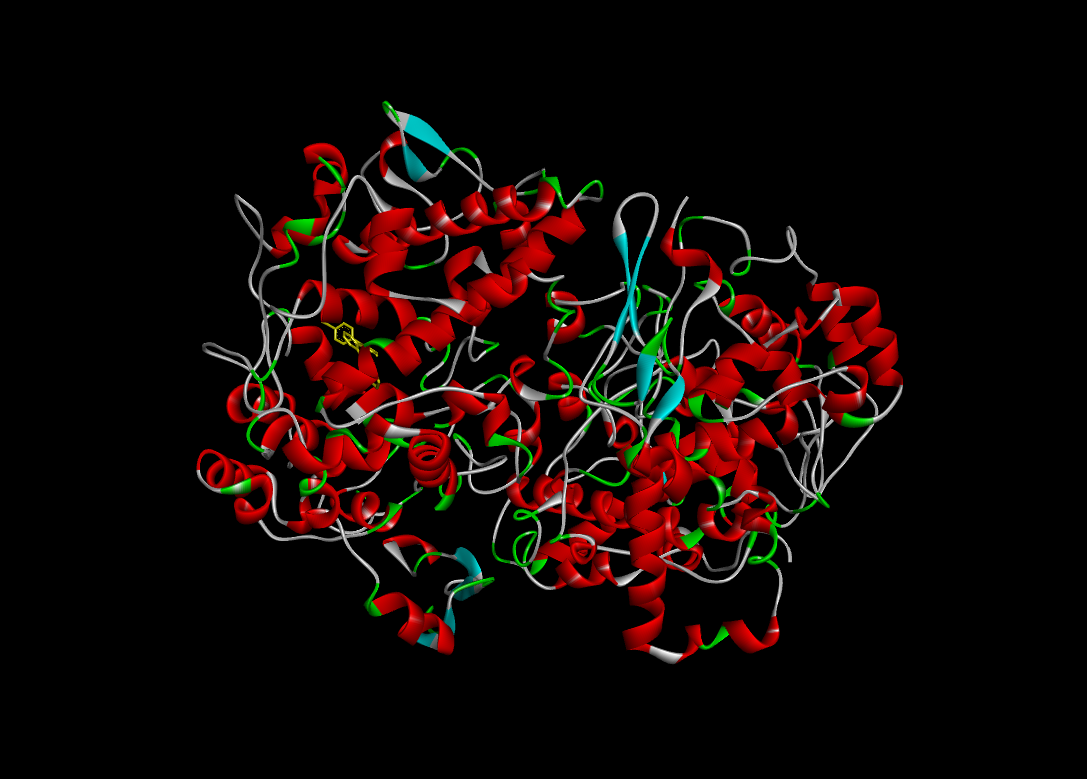
|

|
-11.6 |
|
|
|

|

|
-10.1 |
|
|
|

|
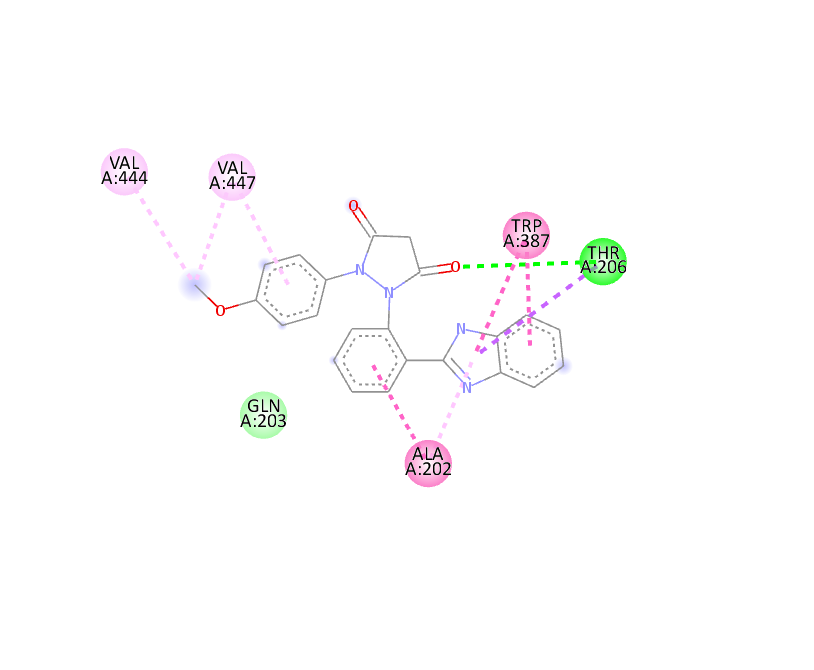
|
-11.5 |
|
|
|
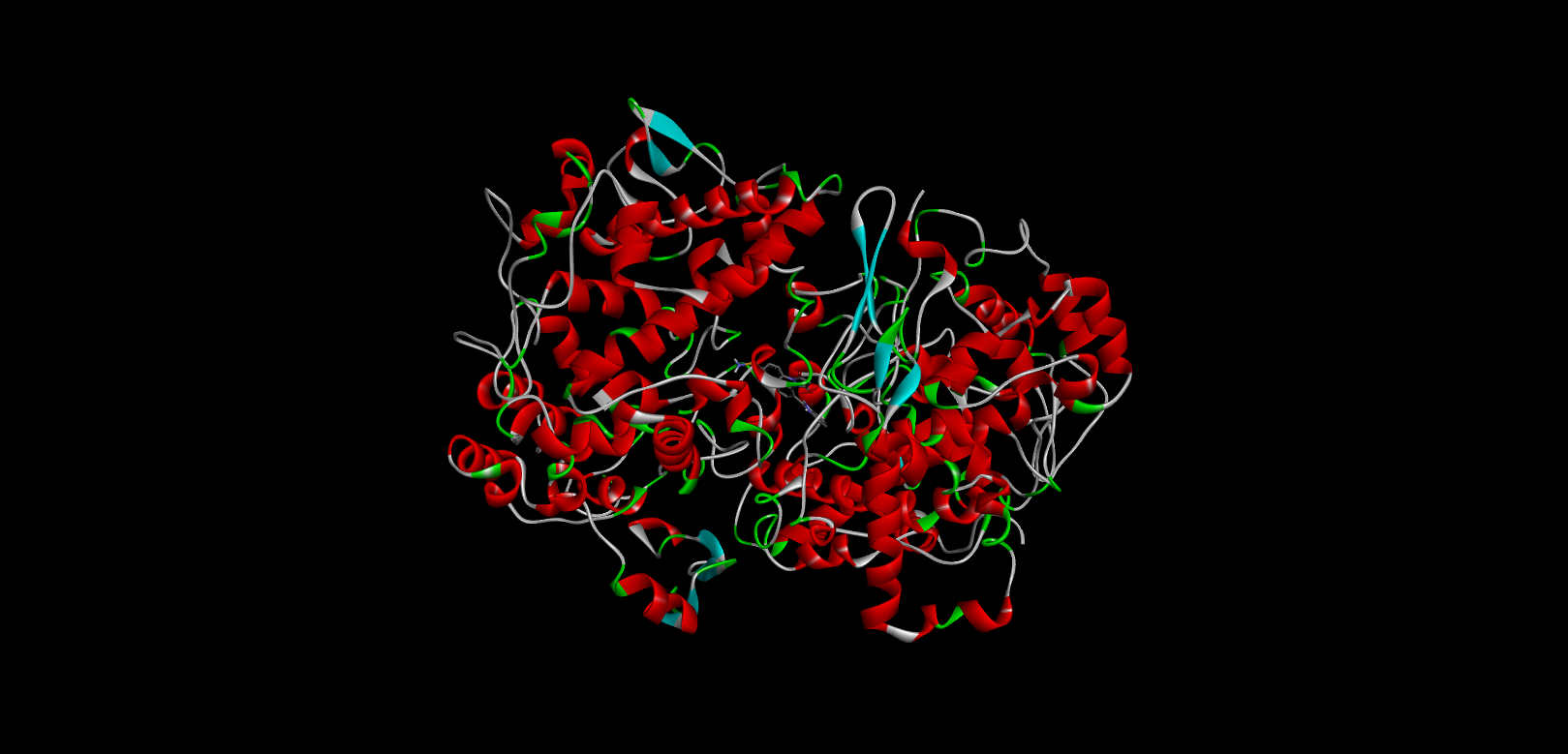
|

|
-11.1 |
|
|
|
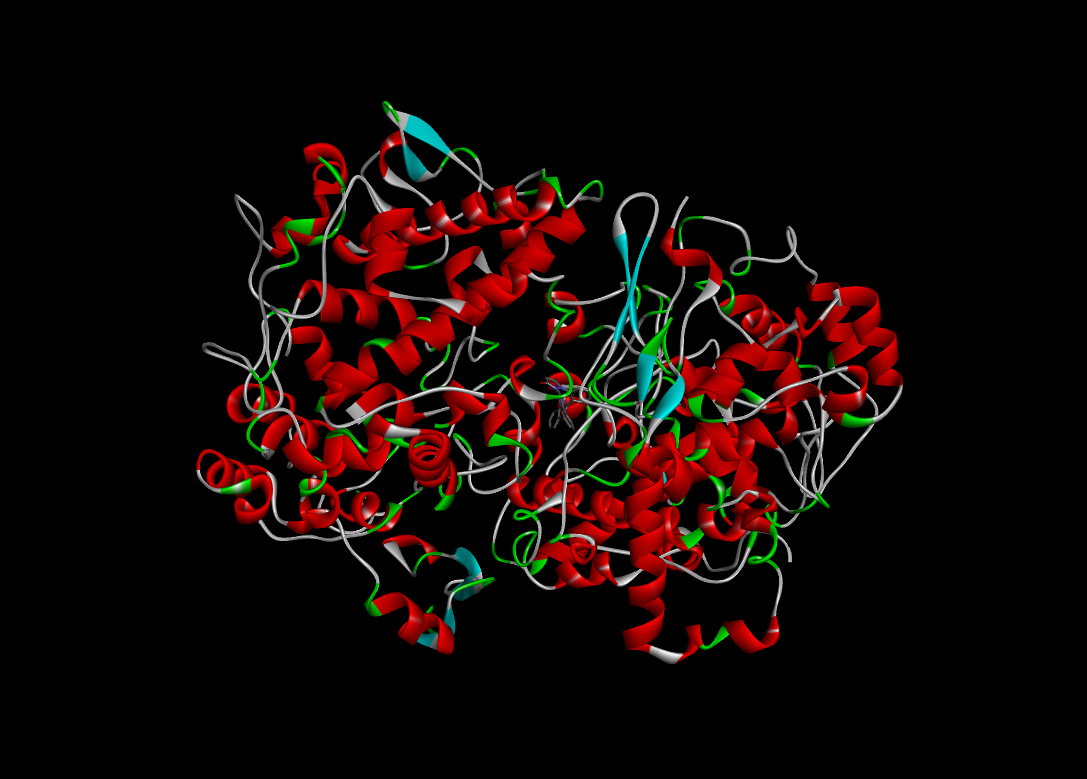
|
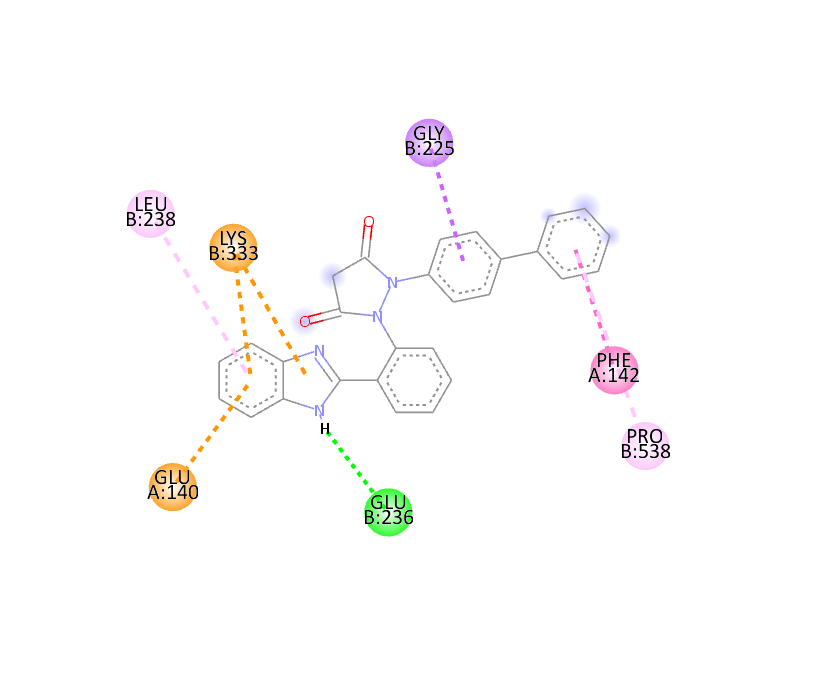
|
-11.6 |
|
|
|
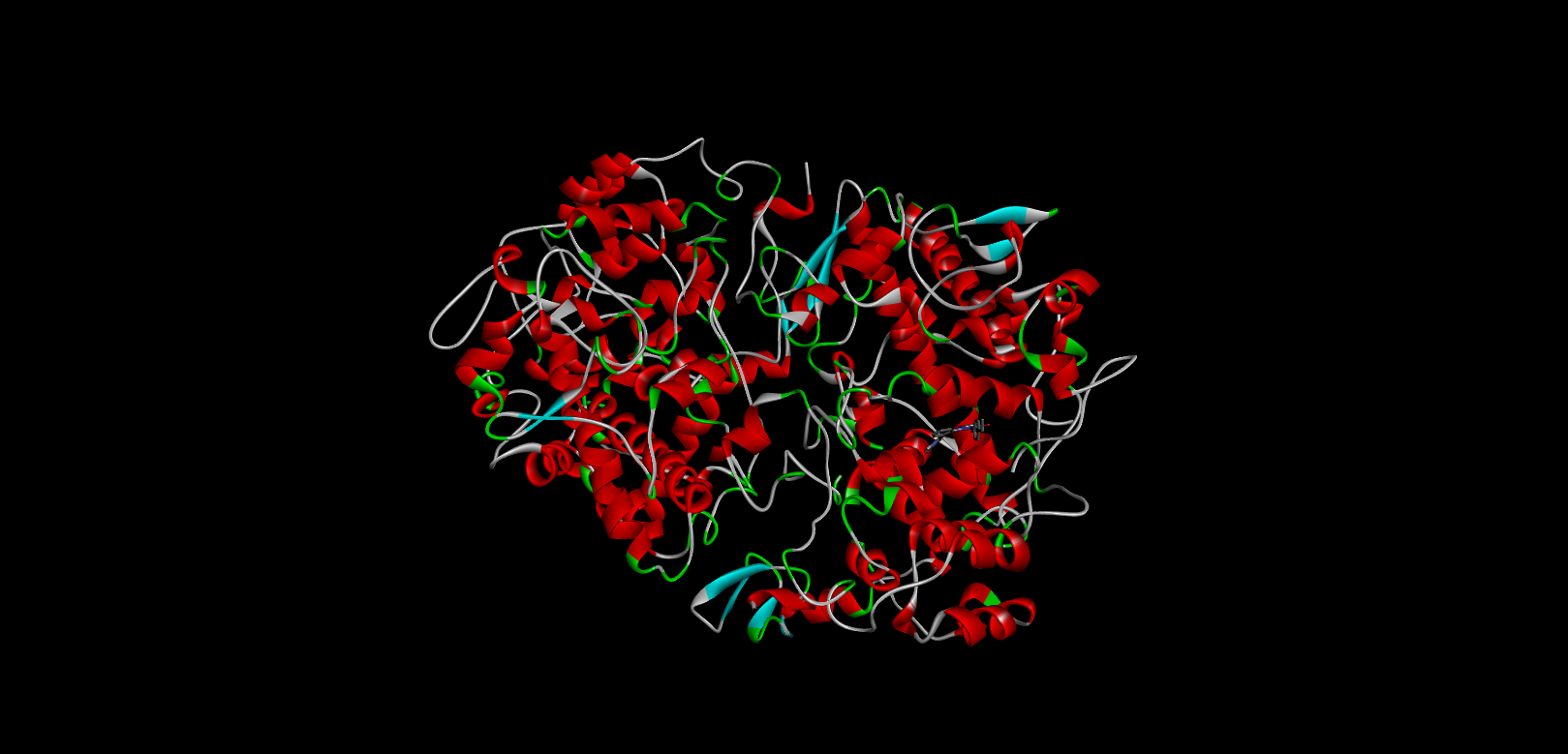
|

|
-11.4 |
CONCLUSION:
In this docking study, the compounds were evaluated Using ArgusLab and PyRx software. The results revealed that these compounds exhibited better binding scores compared to the standard drug, indicating their potential as more effective candidates. These findings suggest that the molecule may possess promising biological activity and ensure further experimental validation.
ACKNOWLEDGEMENT:
The authors wish to thank Sakthi Arul Thiru Amma and Thirumathi Amma ACMEC Trust, for providing facilities to do the working successful manner. We are grateful to thank our Dean Research and Director Academic Prof. Dr. T. Vetrichelvan, M.Pharm., Ph.D., and our principal Dr. D. Nagavalli, M.Pharm., Ph.D., for the kind support and encouraging for the completion of work.
REFERENCES


Sharmila B.*, Dr. G. Abirami, Design and Molecular Docking Analysis of Pyrazolidine-3,5-dione Derivatives as Potential Anti-inflammatory activity Using PyRx and Argus Lab, Int. J. of Pharm. Sci., 2025, Vol 3, Issue 8, 2467-2483 https://doi.org/10.5281/zenodo.16931843
 10.5281/zenodo.16931843
10.5281/zenodo.16931843